Counting Cells
I. INTRODUCTIONCell culture is the technique of growing cell lines or dissociated cells extracted from normal tissues or tumors. The cell count measures the status of the culture at a given time and is essential when subculturing or assessing the effects of experimental treatments on cells. The cell count can be expressed as the number of cells per milliliter of medium (for suspension cultures) or per centimeter squared area of attachment surface (for monolayer cultures).
II. MATERIALS AND INSTRUMENTATION
Dulbecco's phosphate-buffered salire (PBS; DPBS) and exclusion dyes are available from ATCC (DPBS, Cat. No. 30-2200; 0.1% erythrosin B stain solution, Cat. No. 30-2404; 0.4% trypan blue stain solution, Cat. No. 30- 2402). Citric acid and crystal violet are available from Sigma Chemical Co. (citric acid, Cat. No. C-0759; crystal violet, Cat. No. C-0775).
The hemocytometer set and tally counter can be obtained from VWR (double Neubauer chamber and cover glass, Cat. No. 15170-173 or Bright-Line chamber with Neubauer ruling and cover glass, Cat. No. 15170-172; tally counter, Cat. No. 15173-252). The Innovatis "Cedex" cell analysis system and associated reagents are available within the United States from Biomedical Resources International, Inc. (Cedex System, Cat. No. 702 00 00 00; 20-position automatic sampler, Cat. No. AS20, autosampler cups, Cat. No. 600 05 00 00; detergent concentrate, Cat. No. 702.00.125 or 702.00.250; density reference beads, Cat. No. 7 60 00 00 17). The Vi-CELL cell viability analyzer is available from Beckman Coulter (1-position carousel model, Part No. 6605617; 12-position carousel model, Part No. 6605587). The NucleoCounter mammalian cell counting system is available from New Brunswick Scientific (NucleoCounter, Cat. No. M1293-0000; NucleoCassettes, Cat. No. M1293-0100; starter kit, Cat. No. M1293-0020).
III. PROCEDURES
Cell culture measurements can be divided into three major categories: (1) visual methods using light microscopy and a hemocytometer; (2) indirect visual methods, which count cell nuclei using light microscopy and a hemocytometer; and (3) electronic systems, which automate the visual method.
A. Visual Methods
This procedure is taken from those of Absher (1973), Freshney (2000), Merchant et al. (1964), and Phillips (1973).
The most practical method for routine determination of the number of cells that are in suspension or that can be put into suspension is a direct count in a hemocytometer. The hemocytometer is a modified glass slide engraved with two counting chambers of known area. Each chamber grid is composed of nine squares (subgrids), with each square being 1 mm2 (see Fig. 1). The hemocytometer is supplied with a glass coverslip of precise thickness that is supported 0.100mm above the ruled area.
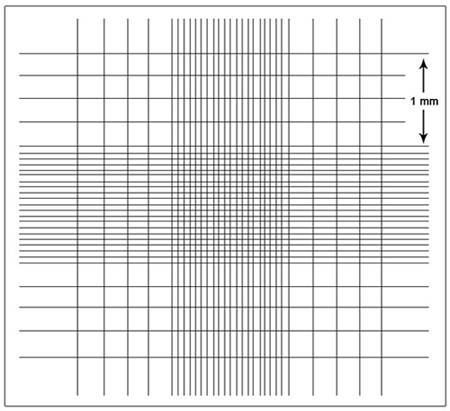 |
| FIGURE 1 Schematic of one chamber grid of a hemocytometer with Neubauer ruling. |
When the coverslip is positioned correctly, the volume of liquid over each square (subgrid) is a known constant, 0.1mm3 or 1.0 × 10-4mL. If cells are counted in each of the corner squares and in the central squares of each of the two chamber grids, a total volume of 1 × 10-3mL (10 × 0.1 mm3= 1.0 mm3) will have been examined, and the cells per milliliter (concentration) of the sample added to the hemocytometer can be determined by multiplying the count by 1000. It should be noted that when cells are diluted in stain or buffer during preparation, then multiplication by the appropriate dilution factor is also required to obtain the concentration of the sample. Determination of viability with the hemocytometer method involves the dye exclusion test. This test relies on the ability of living cells to exclude certain stains from crossing the cell membrane. Dead cells are permeable and will take up the stain.
1. Dilution of Cell Suspension
- Place 0.3 mL of sterile DPBS into a polypropylene tube capable of holding at least a 1.0-mL volume.
- Aseptically collect a sample from the suspension culture or detached monolayer culture. Prior to collection, mix the cell culture to be counted by pipetting up and down two to five times.
- Transfer 0.1 mL of the mixed culture sample to the tube containing the DPBS (Step 1a). Mix by pipetting up and down two to five times.
- Add 0.1 mL exclusion dye (0.1% erythrosin B or 0.4% trypan blue) to the tube containing the cell suspension (step 1c). Mix by pipetting up and down two to five times.
2. Use of Hemocytometer and Counting of Cells
- Obtain a hemocytometer and coverslip from storage (70% alcohol bath). Wipe both pieces dry and put coverslip in place. With a Pasteur pipette or pipetman and pipette tip, introduce enough homogeneous stained cell suspension (Step 1d) to the V-shaped filling troughs for each of the two chambers to fill the hemocytometer.
- Place the hemocytometer on the microscope stage and focus with an objective that will permit one subgrid (one of the nine squares) to be visible (usually 100 × magnification).
- Using a tally counter with at least two registers, count viable (nonstained) cells and nonviable (stained) cells in each grid by counting cells in each of 10 subgrids (5 subgrids per chamber grid). The subgrids to be counted are the 4 outer, corner subgrids and the middle subgrid in each of the two chambers.
- Remove cells from hemocytometer and return it to storage.
3. Calculation of Cell Count
- Calculate the viable cell concentration using the formula C = N × D × 103, where C is the viable cells/mL, N is the number of viable cells counted in 10 subgrids (1.0mm3), D is the dilution factor, and 103 is the hemocytometer correction factor.
- Calculate percentages cell viability using the formula V = N/T × 100%, where V is the percentage viability, N is the number of viable cells counted per 10 subgrids (1 mm3), and T is the number of total cells counted per 10 subgrids (1 mm3).
B. Indirect Visual Methods
This procedure is modified from that of Patterson (1979).
Some adherent cells, such as human diploid cells, are difficult to count directly. In such cases, a procedure can be used to stain and count cell nuclei rather than the cells themselves.
1. Preparation of Solutions
- Prepare 0.1M citric acid by dissolving 1.9212 g in 100 mL distilled water.
- Prepare 0.1M citric acid containing 0.01% (w/v) crystal violet by dissolving 0.005g crystal violet (also known as basic violet 3 or gentian violet; C.I.42555) in 50mL of the 0.1M citric acid (prepared in step Blb)
2. Procedure
- Aseptically transfer a sample from the suspension culture or detached monolayer culture containing approximately 5 × 105 cells to a centrifuge tube. Prior to collection, mix cell culture to be counted by pipetting up and down two to five times.
- Centrifuge at 500 ± 50g for 5 to 10 min.
- Decant the supernatant. Add 1.0mL 0.1M citric acid solution to the cell pellet. Mix well and incubate at 35° to 37°C for 1 to 2 h.
- Separate nuclei by violent shaking followed by centrifugation at 1000 ± 100 g for 20 to 25 min.
- Discard supernatant. Resuspend the cell pellet in 0.5 to 1.0 mL citric acid-crystal violet solution.
3. Use of Hemocytometer and Counting of Cells
- Obtain a hemocytometer and coverslip from storage (70% alcohol bath). Wipe both pieces dry and put coverslip in place. With a Pasteur pipette or pipetman and pipette tip, introduce enough homogeneous stained cell suspension (step 1d) to the V-shaped filling troughs for each of the two chambers to fill the hemocytometer.
- Place the hemocytometer on the microscope stage and focus with an objective that will permit one subgrid (one of the nine squares) to be visible (usually 100X magnification).
- Using a tally counter, count stained nuclei in each grid by counting nuclei in each of 10 subgrids (5 subgrids per chamber grid). The subgrids to be counted are the 4 outer, corner subgrids and the middle subgrid in each of the two chambers.
- Remove cells from hemocytometer and return it to storage.
4. Calculation of Cell Count
Calculate the cell concentration using the formula C = N × D × 103, where C is the cells/mL, N is the number of nuclei counted in 10 subgrids (1.0mm3), D is the dilution factor, and 103 is the hemocytometer correction factor.
C. Electronic Systems The visual method can be user dependent, tedious, and time-consuming. Over the course of the last few years, systems that eliminate these problems through the automation of the process have become available commercially. There are two major types of system: (1) those that use trypan blue to arrive at a viable cell count and (2) those that use propidium iodide (PI) for counting cell nuclei.
1. Automated Trypan Blue Method
Both the Cedex system (Innovatis) and the Vi-CELL system (Beckman Coulter) incorporate image analysis technology and fluidics management to automate the mixing, staining, and hemocytometer counting process. As a result, the staining process is executed with high precision and reproducibility. Automation of the sample preparation process eliminates human errors in connection with pipette handling.
- Set up the system with the appropriate reagents following the manufacturer's instructions.
- Calibrate the system as per the manufacturer's schedule and instructions.
- Aseptically collect a sample from the suspension culture or detached monolayer culture. Prior to collection, mix the cell culture to be counted by pipetting up and down two to five times. Transfer at least 1.0mL to a sample cup.
- Load sample cup onto the unit.
- Log in pertinent information about the sample and initiate the counting sequence.
- When the count cycle is complete (typically about 3 min), the viable cell density, total cell density, and percentage viability will appear on the screen. The results can then be printed and saved as appropriate.
- After use, the sample cup(s) can be disposed of as biological waste. The system flow path is flushed and cleaned automatically, following each count, as part of the count cycle.
2. Automated Propidium Iodide Method
Similar to the Cedex and Vi-CELL systems, the NucleoCounter mammalian cell counting system (New Brunswick Scientific) also incorporates image analysis technology and fluidics management to automate the mixing, staining, and hemocytometer counting process. Unlike these systems, however, the NucleoCounter stains nuclei rather than intact cells. Because of this, two separate counts are needed to obtain culture viability.
- Aseptically collect two samples from the suspension culture or detached monolayer culture. Prior to collection, mix the cell culture to be counted by pipetting up and down two to five times.
- Pretreat the first sample with lysis and stabilization buffers following the manufacturer's instructions. Load the pretreated sample into the NucleoCassette. Place into the unit and press "run."
- A total cell count will be displayed when the run is complete (typically about 30s).
- Load the second, untreated sample into another NucleoCassette. Place into the unit and press "run."
- A total cell count will be displayed when the run is complete (typically about 30s).
- Divide the total count obtained from the pretreated sample (step C2b) by the total count obtained from the untreated sample (step C2d) and multiply by 100% to obtain the percentage viability of the culture.
- After use, the NucleoCassette(s) can be disposed
of as biological waster, with the PI dye safely enclosed.
The system itself requires no cleaning.
IV. PITFALLS
- The visual method is best applied when only a few samples are to be counted. If numerous cultures are to be counted on a routine basis, electronic systems should be considered.
- The dye exclusion test gives information about the integrity of the cell membrane, but does not necessarily indicate how the cell is functioning.
- Trypan blue has a greater affinity for serum proteins than for cellular protein. It is recommended that cells be suspended in DPBS or serum-free medium before counting when using this exclusion dye. When using trypan blue as the exclusion dye, mix the sample well and allow it to stand for 5 to 10 min prior to performing the cell count.
- Among the major errors in all visual counts are:
- Nonuniform cell suspensions. The total volume over a chamber grid is assumed to be a random sample. This assumption is invalid unless the cell suspension is uniform and free of cell clumps. Also, cells settle from suspensions rapidly. A uniform suspension can only be obtained if cells are mixed thoroughly before sampling.
- Improper filling of chamber grids. The chamber must be filled by capillary action, without overflowing. Both pipettes and the hemocytometer set must be scrupulously clean.
- Failure to adopt a routine method for counting cells in contact with boundary lines. All such conventions are arbitrary, but are essential to obtain reproducible results by counting comparable areas. The most commonly used method is to count only those cells that touch the left and/or upper boundary lines. Those cells that touch the right and/or lower boundary lines are not counted.
- Statistical error. The cell sample should be diluted so that no fewer than 10 and no more than 50 cells are over each 1-mm 2 square. Because the cells follow a Poisson distribution in the hemocytometer chamber grids, the count error will be approximated by the square root of the count. Using the best techniques, an experienced individual can generally obtain counts with a total error of 10 to 15% (Absher, 1973).
- The dynamic range for the Cedex and Vi-CELL systems is 1 × 104 to 1 × 107 cells/mL. For the Nucleo- Counter, it is 5 × 103 to 4 × 106, with the optimum being 105 to 2 × 106 cells/mL. Samples should be diluted or concentrated as appropriate and rerun if the values reported by the system fall outside these ranges.
- Due to the small bore of the flow cell path in the Cedex and Vi-CELL systems, routine maintenance of the instruments as outlined by the manufacturer is critical. Failure to perform routing priming and/or flushing could result in clogging.
References
Absher, M. (1973). Hemocytometer counting. In "Tissue Culture: Methods and Applications" (P. E Kruse, Jr., and M. K. Patterson, Jr., eds.), pp. 395-397. Academic Press, New York.
Freshney, R. I. (2000) Quantitation. In "Culture of Animal Cells: A Manual of Basic Technique," 4th Ed., pp. 309-312. Wiley-Liss, New York.
Merchant, D. J., Kahn, R. H., and Murphy, W. H., Jr. (1964). "Handbook of Cell and Organ Culture," 2nd Ed., pp. 155-157. Burgess, Minneapolis.
Patterson, M. K. (1979). Measurement of growth and viability of cells in culture. In "Cell Culture" (W. B. Jakoby and I. H. Pastan, eds.), pp. 141-149, Academic Press, New York.
Phillips, H. J. (1973). Dye exclusion test for cell viability. In "Tissue Culture: Methods and Applications" (P. E Kruse, Jr., and M. K. Patterson, Jr., eds.), pp. 406-408. Academic Press, New York.




