Amino Acids
All proteins are synthesized from the 20 α-amino acids specified by the genetic code as shown in Fig. 1. The nature of an amino acid is determined by the “sidechain” attached to the α-carbon (Table I). All of these amino acids, except for glycine which carries two hydrogens on its α-carbon, have a chiral center located at the α-carbon. Thus the amino acids exist as either the L- or D-isomers. Only the L-stereoisomer is utilized in protein biosynthesis (Fig. 2). This introduces chirality into all protein molecules that is the source of most of the asymmetric features found in protein structures. The use of only one of the two stereoisomers of the amino acids also establishes a structural uniqueness that is essential for biochemical specificity.There are four classes of amino acids specified by the genetic code:
1 aliphatic amino acids,
2. aromatic amino acids,
3. polar amino acids, and
4. charged amino acids.
These groups of amino acids provide the range of properties necessary to create a stable, functional folded protein.
As discussed elsewhere the primary driving force in protein folding and protein structure is the hydrophobic effect. This serves to sequester the hydrophobic side chains away from the bulk solvent. Once folded a typical protein is a densely packed entity that contains few holes larger than a water molecule. The aliphatic amino acids which include glycine, alanine, valine, leucine, isoleucine, and proline provide the range of small hydrophobic amino acids necessary to fill the gaps in the interior of the protein. Glycine and proline serve special roles in protein structure. Glycine is the smallest amino acid and is unique because it lacks a side chain. This gives it more conformational freedom than any other amino acid. Glycine is often found in turns and loops where other amino acidswould be sterically unacceptable. It is also found where secondary structural elements intersect and other side chains would introduce molecular collisions. In contrast proline is unusual because it is conformationally restricted. As such it is often found in turns since it introduces an inherent kink in the polypeptide chain without any entropic cost to protein folding. Proline is also unique in that it is the only amino acid (or technically an “imino acid”) that is commonly found to form a cis peptide bond between itself and the residue that precedes it in the polypeptide chain. In this instance the energy barrier to rotation is considerably less than all other peptide bonds (13 kcal/mol vs ∼20 kcal/mol). This post-translational conformational modification often represents a slow step in protein folding.
Phenylalanine, tyrosine, and tryptophan are large aromatic residues that are normally found buried in the interior of a protein and are important for protein stability. Tyrosine has special properties since its hydroxyl side chain may function as a powerful nucleophile in an enzyme active site (when ionized) and is a common site for phosphorylation in cell signaling cascades. Tryptophan has the largest side chain and is the least common amino acid in proteins. It has spectral properties that make it the best inherent probe for following protein folding and conformational changes associated with biochemical processes.
The polar amino acids include serine, threonine, cysteine, methionine, asparagine, and glutamine. These are an important class of amino acids since they provide many of the functional groups found in proteins. Serine often serves as a nucleophile in many enzyme active sites, and is best known for its role in the serine proteases. Both serine and threonine are sites of phosphorylation and glycosylation which are important for enzyme regulation and cell signaling. Cysteine is the most reactive amino acid side chain. It serves as a potent nucleophile and metal ligand (particularly for iron and zinc), but is best known for its ability to form disulfide bonds, which often make an important contribution to the stability of extracellular proteins. Methionine is a fairly hydrophobic amino acid and typically found buried within the interior of a protein. It can form stacking interactions with the aromatic moieties of tryptophan, phenylalanine, and tyrosine. Asparagine and glutamine are close relatives of aspartate and glutamate but differ in the lack of charge and altered hydrogen bonding characteristics. In general these are not very reactive residues; however, asparagine is a common site for glycosylation.
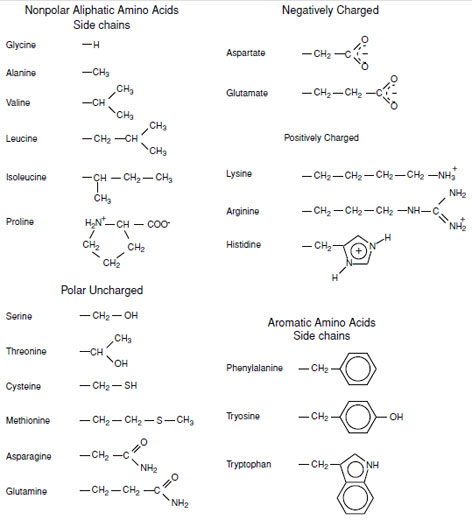 |
| FIGURE 1 The 20 amino acid side chains specified by the genetic code. All except glycine have a β-carbon. Proline is technically an imino acid since it is a secondary amine. |
| Amino acid | Side chain pKa | Occurrence | ||||
|---|---|---|---|---|---|---|
| Aliphatic | ||||||
| Glycine | Gly | G | 7.2 | |||
| Alanine | Ala | A | 7.8 | |||
| Valine | Val | V | 6.6 | |||
| Leucine | Leu | L | 9.1 | |||
| Isoleucine | Ile | I | 5.3 | |||
| Proline | Pro | P | 5.2 | |||
| Aromatic | ||||||
| Phenylalanine | Phe | F | 3.9 | |||
| Tyrosine | Tyr | Y | 10.5 | 3.2 | ||
| Tryptophan | Trp | W | 1.4 | |||
| Polar uncharged | ||||||
| Serine | Ser | S | ∼13 | 6.8 | ||
| Threonine | Thr | T | ∼13 | 5.9 | ||
| Cysteine | Cys | C | 8.4 | 1.9 | ||
| Methionine | Met | M | 2.2 | |||
| Asparagine | Asn | N | 4.3 | |||
| Glutamine | Gln | Q | 4.3 | |||
| Positively charged | ||||||
| Lysine | Lys | K | 10.5 | 5.9 | ||
| Arginine | Arg | R | 12.5 | 5.1 | ||
| Histidine | His | H | 6.0 | 2.3 | ||
| Negatively charged | ||||||
| Aspartate | Asp | D | 3.9 | 5.3 | ||
| Glutamate | Glu | E | 4.1 | 6.3 | ||
Lysine, arginine, and histidine can carry a positive charge. Of these, arginine is constitutively positively charged since its pKa lies around 12.5. Lysine also plays an important role in coordinating negatively charged ligands; however, it functions as a nucleophile in some enzyme catalyzed reactions. Histidine is perhaps the most common and versatile catalytic residue in proteins. Its pKa of ∼6.0 allows it to function both as a catalytic acid or base at physiological pH depending on its local environment. Histidine also has the ability to form covalent intermediates during catalysis such as phosphohistidine. In addition, it is often a ligand for transition metal ions such as iron and zinc.
A. Post-Translational Modifications
Once synthesized and folded, many proteins undergo post-translational modifications before they reach a functional state. Over 200 variant amino acid residues have been identified in proteins thus far. These changes are almost always achieved through an enzymatic pathway. The simplest changes include the formation of disulfide bonds (discussed later) and proteolytic processing of the polypeptide chain to yield a functional protein. Examples of proteolytic processing include the removal of signal peptides, the activation of zymogens to generate active forms of many proteolytic enzymes, and the maturation of viral proteins. Additionally proteolytic processing occurs in the biosynthetic pathway of many hormones. Other simple changes include the glycosylation of asparagine, serine, threonine, and phosphorylation of serine and tyrosine.
It is noteworthy that many post-translation modifications are associated with a sequence motif such that it is frequently possible to identify potential sites directly from the amino acid sequence. This arises because most post-translational modifications are the result of enzymatic pathways, which are usually highly specific. Thus protein sequences inferred from DNA sequence are often annotated with sites for post-translational modification. These sites should be viewed with caution since proof of modification can only be obtained through chemical or physical experimentation.
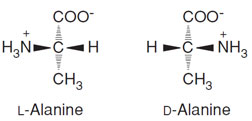 |
| FIGURE 2 Stereoisomers of L-alanine and D-alanine. |
Although many proteins derive all of their function from their constitutive amino acids, a large number of proteins require additional cofactors in order to fulfil their biological role. These cofactors provide chemical properties that are not present in the 20 amino acid residues. For example, none of the amino acids are capable of facilitating an oxidation/reduction reaction. A wide range of cofactors are utilized including inorganic ions such as Fe2+,Mg2+, and Zn2+, or complex organic molecules that are normally described as coenzymes such as flavin adenine dinucleotide or nicotinamide adenine dinucleotide. If the coenzyme is covalently bound to the enzyme it is often called a prosthetic group. The complete enzyme is called a holoenzyme, whereas the protein in the absence of its cofactors is called an apoenzyme or apoprotein. Many apoproteins are considerably less stable in the absence of their cofactor. This suggests that although the amino acid sequence dictates the overall threedimensional structure, the cofactor is an integral part of the protein.
C. Context Determines Function
Typically proteins and especially enzymes contain only a few residues that are absolutely vital for function. In contrast there are usually many other residues of the same type in the protein that do not fulfill any special role. In many cases the catalytic residues have different chemical and physical properties from the same amino acid in solution; for example, the pka of the side chain might be several pH units higher or lower than the free amino acid. It is generally found that the behavior of an amino acid is profoundly influenced by the context of that amino acid within the protein. Altering the chemical properties of a functional group is one of the major attributes of protein structure and appears to be essential for the activity of most enzymes. There are many ways in which this is achieved; however, a simple example is placement of a charged residue in the interior of a protein such that the deionized state is favored. This serves to raise the pka of aspartate and glutamate and lower the pka of lysine.




