Folding Pathways
The intramolecular interactions discussed above stabilize the final folded structure of a protein. However, knowledge of the end states, N and U, tells us nothing of the path taken between them. Proteins fold on the time scale of microseconds to hundreds of seconds. It is impossible to sample all possible conformations during this time and it is clear that there is a preferred order of events leading to the final tertiary fold. Determining this order of events is an area of active inquiry. The questions that experimentalists are attempting to answer are “Do autonomously folding substructures nucleate the folding of other regions of the protein?” or “Do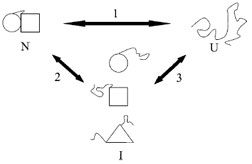 |
| Figure 2 Illustration of cooperative vs. noncooperative unfolding transitions. If the native state of a protein (N) is denatured inTo = ΔSo the unfolded state (U) in a single transition (pathway 1), then it is a two-state or cooperative unfolding transition. Alternatively, the native state may be converted inTo = ΔSo one or more intermediate states (pathway 2). For example, if a protein is comprised of multiple domains, one of the domains may be unfolded first. It is also possible to form a completely different intermediate before Unfolding completely. The presence of intermediate species may be observed using kinetic or equilibrium techniques. However, intermediates detectable by kinetic methods may or may not be observable by equilibrium methods. |
It is clear that the kinetics of protein folding is protein dependent. Some fold in a distinctly cooperative fashion, such that one can detect only the unfolded and native end states (U ↔ N), being two-state in a kinetic as well as equilibrium sense. This is equivalent to saying that there is a single rate-limiting step, and intermediate species are not populated. Alternatively, some proteins fold by populating one or more distinct intermediate species (e.g., U ↔ I ↔ N; see Fig. 2). Thus, formation of the intermediate species is fast, often formed in the dead-time of the instrument, and formation of the native species from the intermediate is relatively slow and easily monitored experimentally. It has been shown that this slow phase in some cases may be due to proline isomerization.




