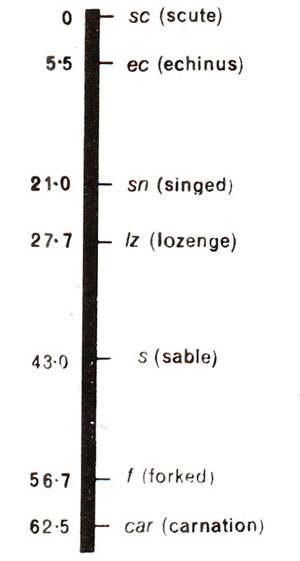
Fig. 10.11. A part of linkage of X-chromosome in Drosophila.
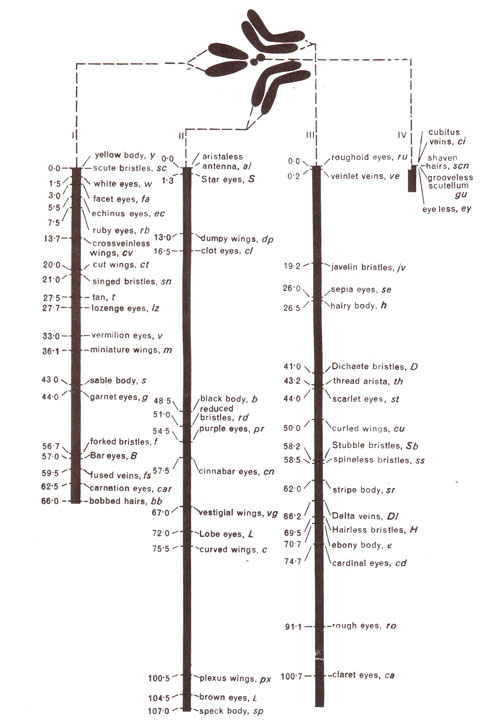
Fig. 10.12. Linkage maps of four different chromosomes of Drosophila.
Linkage maps have been prepared in large number of animals and plants with the help of recombination frequencies as discussed above. By testing different groups of genes having atleast one gene in common, additional genes can be added in a linkage map. It is important to notice that recombination frequencies and map distances do not correspond, because over longer distances, although the map distance will exceed even 100, but recombination frequency between any two genes will never exceed 50, as earlier discussed, since double and multiple crossovers will not be discovered. For illustration, a part of linkage-map of X-chromosomes in
Drosophila, as initially worked out, is given is Figure 10.11. The map distances are given by recombination frequencies and are sometimes represented in map units (m.u.) also called centi Morgan (1 centi Morgan = 1 cM = 1% recombination).
Linkage maps of four different chromosomes of
Drosophila are shown in Figure 10.12. X-chromosome has 66 crossover units, with genes
yellow and
bobbed at two extreme ends of the map. But this does not mean that recombination frequency between
yellow and
bobbed will be 66 units. In fact, we know that recombination frequency will never exceed 50% between any two loci. These 66 units will be actually obtained by making use of a
mapping function.
It is thus evident that we should not confuse
map distances with recombination frequencies.
Based on data from
Drosophila, corn and
Neurospora it became clear that relationship between map distances and recombination frequencies assumes a curve shown in Figure 10.15. The curve suggests that at lower values there is a linear relationship, but as recombination value approaches towards 50%, the linear relationship is gradually lost and recombination frequency is always less than the map distance, so much so that at 50% recombination, further increase in map distance does not cause any increase in recombination value.

Fig. 10.11. A part of linkage of X-chromosome in Drosophila.

Fig. 10.12. Linkage maps of four different chromosomes of Drosophila.






