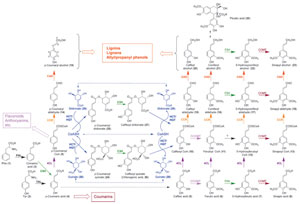
Biosynthesis of Monolignols
The phenylpropanoid pathway (Fig. 13.1) entry point is generally considered to be the amino acid phenylalanine (1, Phe), through action of phenylalanine ammonia lyase (PAL, EC 4.3.1.5) forming trans-cinnamic acid (3). Some monocots (e.g., maize), though, are also able to utilize the p-hydroxylated amino acid tyrosine (2, Tyr) as a partial entry point to this pathway through tyrosine ammonia lyase (TAL). The mechanism of phenylalanine (1) formation in planta has been a matter of debate, with arogenate decarboxylation/dehydration appearing to be the preferred biosynthetic route (Cho et al., 2007; Jung et al., 1986). The subsequent deamination reaction, catalyzed by PAL [forming cinnamic acid (3) and stoichiometric amounts of ammonium ion], was discovered by Eric Conn and his group (Koukol and Conn, 1961) and is probably the most extensively studied reaction in plant secondary metabolism (Lewis et al., 1999).The aromatic amino acid ammonia lyases (PAL, TAL, and HAL, histidine ammonia lyase) all possess the interesting internal cofactor 3,5-dihydro-5-methylidene-4H-imidazol-4-one (MIO) formed by spontaneous dehydration and cyclization of a conserved tripeptide sequence (Ala-Ser-Gly) (Baedeker and Schulz, 2002; Calabrese et al., 2004; Schwede et al., 1999), which acts as a strong electrophile during catalysis. The mechanism of ammonia elimination is still controversial, with the two most likely possibilities currently being (1) a E1cb-like mechanism where MIO forms a bond to the amine group of the amino acid (Fig. 13.3A) or (2) a Friedel-Crafts-like mechanism where MIO reacts with the aromatic ring of the substrate, which therefore loses aromaticity and increases the acidity of the nearby β-proton (Fig. 13.3B) (Calabrese et al., 2004; Louie et al., 2006; Poppe and Rétey, 2005). Recent studies of TAL from Rhodobacter sphaeroides have determined
that Tyr (2) preference (i.e., TAL activity) is largely dictated by one specific TAL-conserved active site residue, H89, which is thought to be hydrogenbonded to the phenolic moiety of the substrate Tyr (2). When H89 is replaced with a Phe residue (characteristic of PAL), substrate selectivity is switched and the mutant H89F TAL prefers Phe (1) (i.e., PAL activity). When the corresponding inverse mutation was made in PAL from Arabidopsis thaliana, the mutant (F144H) PAL enzyme lost PAL activity and instead gained TAL activity (Louie et al., 2006; Watts et al., 2006).
Although initially regarded as a rate-determining reaction in the pathway (Camm and Towers, 1973; Rubery and Fosket, 1969), PAL does not seem to serve as a major carbon allocation regulator, with flux apparently being determined by the availability of Phe (1) and by downstream enzymes in the pathway, especially C4H/C3H (Anterola et al., 1999, 2002). In A. thaliana, a four-membered PAL multigene family (AtPAL1–4) has been characterized, with Tyr (2) serving, as expected, as a very poor substrate. Kinetic parameters for AtPAL1, 2, and 4 with Phe (1) were similar, with Km values of ~60–70 µM and V
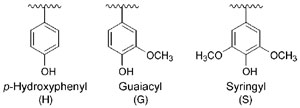 |
| FIGURE 13.2 Aromatic (H, G, and S) residues in plant lignins. |
max ~5–10 pkat/µg protein, whereas AtPAL3 was barely active (Km = 2.56 mM, Vmax = 0.4 pkat/µg protein) (Cochrane et al., 2004). Rohde et al. (2004) have also analyzed the effects of PAL downregulation in A. thaliana. In particular, pal1 and pal2 mutants apparently presented no clear phenotypic alterations, with lignin levels of approximately 60–70% of wild type (WT), perhaps as a result of compensatory upregulation of the remaining PAL genes in each mutant (i.e., AtPAL2/4 were upregulated in pal1 mutants, AtPAL1/4 in pal2 mutants). Double pal1 pal2 mutants, on the other hand, exhibited approximately 25% of WT PAL activity (resulting mainly from the presence of AtPAL4), were sterile, and had estimated lignin levels of approximately 30–35% of WT, with this lignin having its S/G ratio apparently nearly doubled relative to WT.
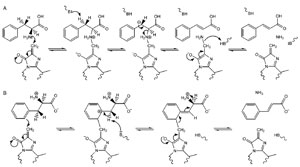 |
| FIGURE 13.3 Potential mechanisms for PAL catalysis: (A) E1cb-like mechanism and (B) Friedel-Crafts-like mechanism. Source: Redrawn from Calabrese et al. (2004). |
Trans-cinnamic acid (3) is next para-hydroxylated by cinnamate 4-hydroxylase (C4H, EC 1.14.13.11), a membrane-associated cytochrome p-450 hydroxylase acting along with an associated NADPH-dependent reductase (Lewis et al., 1999), to afford p-coumaric acid (4). Subsequent ring hydroxylation reactions [i.e., C3H-catalyzed 3-hydroxylation leading to formation of the catechol-like substructure of caffeic acid (5) and derivatives, as well as F5H-catalyzed 5-hydroxylation leading to the 5-hydroxyguaiacyl substructure present in 5-hydroxyconiferyl alcohol (22)] are also performed by cytochrome p-450 hydroxylases. Therefore, these enzymes (C4H, C3H, and F5H) are often grouped together as a single class. In A. thaliana, each of these enzymes is encoded by either one- (C4H and C3H) or two-membered (F5H) gene families, with locus numbers At2g30490 (C4H), At2g40890 (C3H), and At4g36220/At5g04330 (F5H), respectively. As of yet, no X-ray crystal structures for any of these plant cytochrome p-450 enzymes have been reported, although they play important (i.e., possibly regulatory, in the case of C4H and C3H) roles in monolignol biosynthesis and lignification.
Interestingly, 3-hydroxylation of the p-coumaric acid (4) moiety apparently occurs only after esterification, first to its CoA form (see below), then to shikimate and/or quinate esters; that is, p-coumarate 3-hydroxylase (C3H) acts on p-coumaroyl quinate (24) and/or shikimate (25) to form the corresponding caffeic acid derivative (26 and/or 27, respectively). Recognition that the p-coumaroyl esters serve as substrates for cytochrome p-450 hydroxylation was established using parsley (Petroselinum crispum) cell suspension cultures (Heller and Kühnl, 1985); more recent studies identified the gene and expressed functional recombinant protein (Schoch et al., 2001). In this regard, transesterases (HCT/HQT) that transfer hydroxycinnamoyl groups from the corresponding CoA esters to shikimate (29) and quinate (28), respectively, were also recently reported (Hoffmann et al., 2003; Niggeweg et al., 2004). HCT, which prefers shikimate (29) as an acyl acceptor, has been reported to be present in stem vascular tissues in tobacco, silencing of which in A. thaliana led to dwarfed plants. In HCT-silenced Nicotiana benthamiana plants, the corresponding lignin reportedly decreased and presented lower S and increased H contents, respectively, that is, in agreement with its presumed participation in monolignol biosynthesis (Hoffmann et al., 2004).
The final ring hydroxylation catalyzed by the cytochrome p-450 F5H was first reported to act on ferulic acid (6) (Grand, 1984); however, findings by Fukushima and colleagues indicated that the hydroxylation step preferentially involved coniferyl alcohol (21) (Chen et al., 1999a,b). The corresponding F5H (CYP84) gene was subsequently isolated from A. thaliana (Meyer et al., 1996), and further analysis of the recombinant CYP84 protein indicated a preference for both coniferyl alcohol (21) and coniferyl aldehyde (16) rather than ferulic acid (6) as substrates [Km values of 3 and 1 µM, Vmax of 6 and 5 pkat/mg protein for 21 and 16, respectively, while Km = 1 mM and Vmax = 4 pkat/mg protein for the latter (6)] (Humphreys et al., 1999), which is in agreement with Fukushima’s seminal findings.
Through further modification of the aromatic ring, the catechol-like substructure of caffeic acid (5) [or an ester derivative thereof, e.g., caffeoyl CoA (10)], caffeyl aldehyde (15), and/or caffeyl alcohol (20) can serve as substrate for SAM (S-adenosyl methionine)-dependent O-methyltransferases (OMTs). In this way, the guaiacol substructure of ferulic acid (6) [or the respective ester derivative, e.g., feruloyl CoA (11)], coniferyl aldehyde (16), and/or coniferyl alcohol (21), respectively, can be formed. This O-methylation was originally reported to be performed by the so-called caffeic acid O-methyltransferases (COMTs) but was later shown to much more likely utilize caffeoyl CoA (10) as substrate i.e., caffeoyl CoA O–methyltransferase (CCOMT) (Pakusch et al., 1989; Schmitt et al., 1991; Ye et al., 1994). Analogously, the 5-hydroxyguaiacol substructure resulting from F5H-mediated hydroxylation is O-methylated to afford a syringyl moiety. This is catalyzed by an OMT originally thought to act on caffeic acid (5) as primary substrate (COMT), and although the enzyme has now been established to be a 5-hydroxyguaiacyl O-methyltransferase (Anterola and Lewis, 2002; Atanassova et al., 1995), the original ‘‘COMT’’ denomination still largely remains in use.
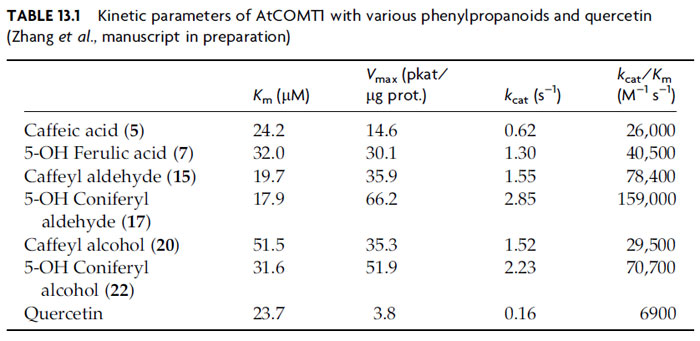
In A. thaliana, 22 genes have been putatively annotated as either COMT (EC 2.1.1.68, 17 putative genes) or CCOMT (EC 2.1.1.104, 5 putative genes). Biochemical analysis of 12 putative AtCOMT proteins bearing highest homology to the previously characterized AtCOMT1 indicated that only the latter is able to efficiently O-methylate a variety of potential substrates, with the remaining 11 not able to act on any of the phenolic compounds tested (Zhang et al., manuscript in preparation). Caffeyl (15) and 5-hydroxyconiferyl (17) aldehydes are the preferred substrates, having kcat/Km values of 78,400 and 159,000 M-1 s-1, respectively (Table 13.1). This is in accordance with previous analyses (see Anterola and Lewis, 2002), where the largest effect of COMT downregulation was the decrease of S contents in lignin (whereas G levels and total lignin amounts apparently remained relatively unchanged), with the concomitant incorporation of 5-hydroxyconiferyl alcohol (22) into the mutated lignin resulting in weakened and discolored stems. Interestingly, AtCOMT1 has also been described as an enzyme involved in quercetin to isorhamnetin formation (Muzac et al., 2000); however, the kinetic data indicate that quercetin is a poorer substrate by an order of magnitude (Table 13.1) (Zhang et al., manuscript in preparation).
 |
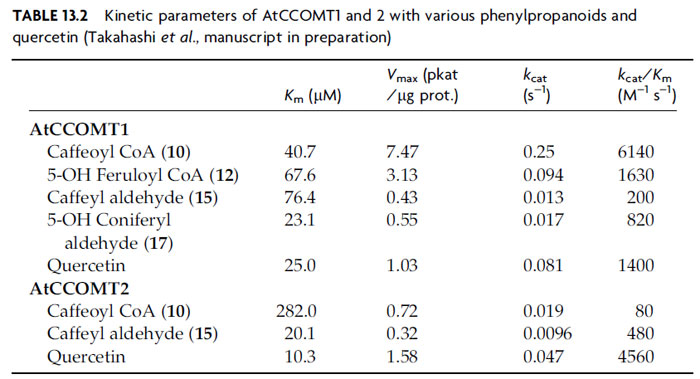
Correspondingly, work on the five putative AtCCOMT enzymes has thus far focused on AtCCOMT1 and 2. AtCCOMT1 uses caffeoyl CoA (10) and 5-hydroxyferuloyl CoA (12) as preferred substrates (Table 13.2), together with quercetin, although with efficacies 10–100-fold lower than those observed for AtCOMT1 acting on the corresponding substrates [caffeyl aldehyde (15) and 5-hydroxyconiferyl aldehyde (17), respectively]. The corresponding acids, as expected, did not serve efficiently as substrates. AtCCOMT2, by contrast, more efficiently methylated quercetin, rather than the possible monolignol pathway intermediates, thus suggesting a role in flavonoid metabolism. Interestingly, AtCCOMT2-catalyzed methylation occurred at both positions 4 and 5 with 5-hydroxyferuloyl CoA (12) and 5-hydroxyconiferyl aldehyde (17) (Takahashi et al., manuscript in preparation), indicating that it has no role in monolignol formation, where position 4 remains unmodified. In summary, while the participation of AtCOMT1 in monolignol biosynthesis in A. thaliana appears to be quite well established (i.e., S monomer formation), the particular roles of the multiple AtCCOMT isoforms, and the particular substrates they act upon, are still a matter of debate and thus remain to be further determined.
 |
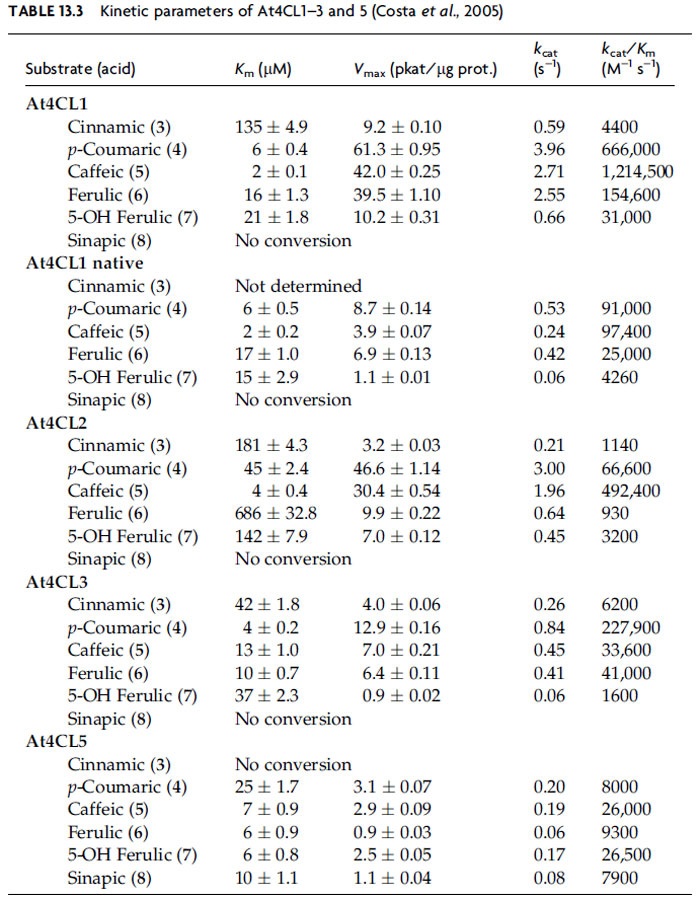
The transformations occurring on the phenolic ring (e.g., hydroxylations, O-methylations) are accompanied by side-chain modification of the aforementioned cinnamic acid derivatives (4–8), first generating CoA esters (9–13) through the twostep activity of hydroxycinnamoyl CoA ligases (4CL, EC 6.2.1.12). These utilize ATP to form an AMp-acyl intermediate that stays enzyme-bound, with AMP then enzymatically displaced by CoA. In A. thaliana, 14 genes had been originally annotated as putative 4CLs, but through further analysis and biochemical characterization, only 4 genes were shown to encode functional 4CL enzymes, albeit with very distinct kinetic properties [using all six possible substrates, cinnamic (3), p-coumaric (4), caffeic (5), ferulic (6), 5-hydroxyferulic (7), and sinapic (8) acids]. At4CL1 is by far the most active, with kcat/Km being ~15–600 times higher than that for others (Table 13.3) (Costa et al., 2005).
 |
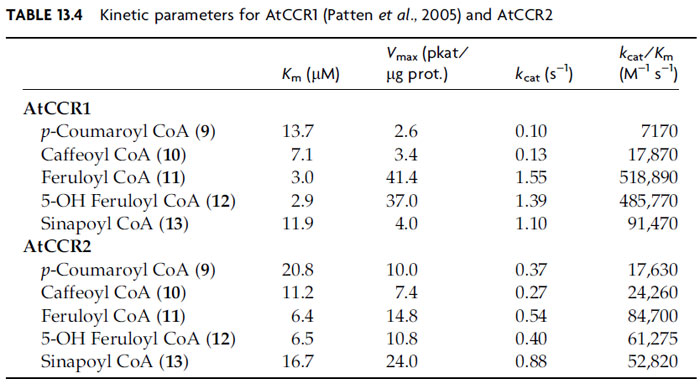
The hydroxycinnamoyl CoA esters (9–13) thus formed serve as substrates for reduction by NADPH-dependent cinnamoyl CoA oxidoreductases (CCR, EC 1.2.1.44) generating the corresponding aldehydes (14–18), with 11 genes being putatively annotated as CCRs in A. thaliana. To date, only three of the putative AtCCRs have been biochemically characterized, namely, AtCCR1, AtCCR2 (Lauvergeat et al., 2001; Patten et al., 2005), and AtCCR8. The most active of these is AtCCR1, which most effectively utilized feruloyl CoA (11) and 5-hydroxyferuloyl CoA (12) as substrates (Table 13.4) (Patten et al., 2005). This is in harmony with the corresponding aldehydes [coniferyl (16) and 5-hydroxyconiferyl (17) aldehydes] serving as substrates for subsequent monolignol formation. AtCCR2, by contrast, was a less effective catalyst, but again slightly preferred compounds 11/12 as substrates (Table 13.4). Interestingly, AtCCR8 displays both CCR and CAD catalytic properties, being able to slowly convert cinnamyl, p-coumaryl (14), caffeyl (15), and sinapyl (18) aldehydes into the corresponding monolignols; however, the physiological significance of these latter observations is still unclear, given the substrate versatility of many proteins.
 |
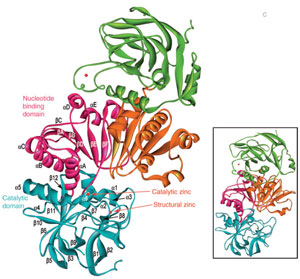
The hydroxycinnamyl aldehydes (14–18) are then further reduced (by cinnamyl alcohol dehydrogenases, CAD, EC 1.1.1.195) to afford the monolignols 19–23 (Lewis et al., 1999). These NADPH-dependent CADs thus catalyze the final step in monolignol biosynthesis and have been studied in some detail in A. thaliana and other plants. In A. thaliana, in silico analyses had indicated the existence of a 17-membered CAD family, but further biochemical examination established that 2 of those (AtCAD4/5) were catalytically the most active (Kim et al., 2004). Moreover, none of the isoforms displayed any strong sinapyl aldehyde (18) preference, contrary to reports of a putative sinapyl alcohol dehydrogenase (SAD) specific for sinapyl aldehyde (18) (Li et al., 2001). X-ray crystal structures for AtCAD4 and 5 have been recently determined (Fig. 13.4), with each being a dimer containing two zinc ions per subunit, one participating in catalysis and the other playing a structural role (Youn et al., 2006a). An A. thaliana cad4 cad5 double mutant has also been obtained (Jourdes et al., 2007; Sibout et al., 2005) and presents a visible prostrate phenotype (i.e., cannot stand upright, see Fig. 13.5A) after approximately 4 weeks growth, resulting from the large decrease in amounts of lignin proper. Furthermore, this double mutant produces only about approximately 10% of WT lignin amounts, based on monolignol-derived thioacidolysis cleavage data, with small amounts of a poly-p-hydroxycinnamyl aldehyde also being formed. Interestingly, deposition of the latter is prematurely aborted relative to lignin proper (Jourdes et al., 2007). This further emphasizes the stringency of the lignification process, where three of the CAD monolignol products (19, 21, and 23) have evolutionarily become the highly conserved substrates for subsequent controlled one electron (radical) oxidation and polymerization of the H, G, and S ‘‘building blocks’’ of lignin (Fig. 13.2) (Anterola and Lewis, 2002). That is, strong evolutionary pressure resulted in lignins being formed from monolignols (as well as, to a much lesser extent, monolignol esters in grasses).