Ion Transport
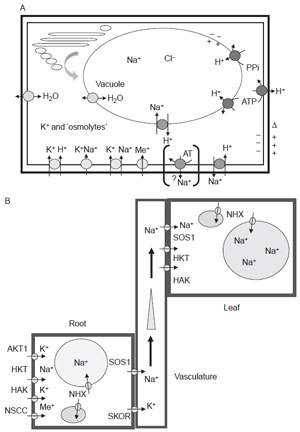
Loss- or gain-of-function experimentation has identified many determinants that control and mediate Na+ and K+ uptake, as well as homeostasis in planta (Amtmann and Sanders, 1999; Hasegawa et al., 2000b; Horie and Schroeder, 2004; Maathuis et al., 1996; Mäser et al., 2002; Niu et al., 1995; Schachtman, 2000; Sondergaard et al., 2004; Tester and Davenport, 2003; Zhu, 2003). Illustrated in Fig. 12.1A are known or suspected cellular Na+ uptake systems, as well as other facilitators of homeostasis such as aquaporins and the proton transporters that establish membrane potentials in different compartments (Gaxiola et al., 2002). Included are channels and transporters responsible for K+ and Na+ flux and H+ pumps that generate the requisite electrochemical potential necessary to facilitate channel function or secondary-active transport (Arango et al., 2003; Borsani et al., 2001; Horie and Schroeder, 2004; Schachtman, 2000; Tester and Davenport, 2003; Vitart et al., 2001; Ward et al., 2003; Zhu, 2003). Other,yet unidentified, genetic loci are involved in the control of Na+ and K+ homeostasis, including those that regulate Ca2+ homeostasis (Nublat et al., 2001; Rus et al., 2001). Focal are Na+ transport systems that control not only intracellular distribution of Na+ but also homeostasis between tissues and organs (Fig. 12.1B) (Berthomieu et al., 2003; Horie and Schroeder, 2004; Laurie et al., 2002; Rus et al., 2004; Tester and Davenport, 2003; Ward et al., 2003; Yokoi et al., 2002). Possibly most important is the salt overly sensitive (sos) pathway described from Arabidopsis, described in detail below (Aharon et al., 2003; Gaxiola et al., 2001; Quintero et al., 2002; Su et al., 2001; Talke et al., 2003). One of the three proteins in this pathway, sos1, controls Na+ uptake into the root xylem. It is hypothesized that high-affinity K+ transporter 1 (HKT1), originally described as a K+ transporter, acts as a Na+ transporter that recirculates the ion from the shoot to the root when Na+ is in excess (Horie and Schroeder, 2004; Munns, 2002; Rus et al., 2004; Shi et al., 2003; Yokoi et al., 2002). Phenotypic suppression of sos1–1 NaCl sensitivity by dysfunctional hkt1 alleles is genetic evidence that these two transport systems have opposing functions in Na+ homeostasis (Laurie et al., 2002). Thus, sos1 and HKT1 may function in concert to regulate Na+ content in the shoot. However, AtHKT1 overexpression does not increase salt tolerance of Arabidopsis as would be expected (Laurie et al., 2002). Most likely the endomembrane cation/H+ transporters, exemplified by the NHX family, are principally responsible for vacuolar compartmentalization of Na+ in plant cells (Aharon et al., 2003; Munns, 2002; Rigas et al., 2001). However, NHX gene copy number and their expression under nonsaline conditions might indicate that the various NHX forms could have very different, additional functions or that their function under saline conditions becomes modified.
Na+ detrimentally affects K+ acquisition. K+ is an essential macronutrient that functions in critical processes ranging from charge balance, osmotic adjustment, and enzyme catalysis to growth and development (Elumalai et al., 2002; Maathuis et al., 1996; Rains and Epstein, 1965). Na+ disturbs intracellular K+ homeostasis possibly because it can compete for binding sites on enzymes and transport proteins (Cramer et al., 1987; Epstein, 1961; Niu et al., 1995; Tester and Davenport, 2003). Ca2+ enhances K+ /Na+ selective accumulation because it facilitates K+ uptake (Epstein, 1961; Hasegawa et al., 2000b; Li et al., 1998).