Ribulose Biphosphate Carboxylase
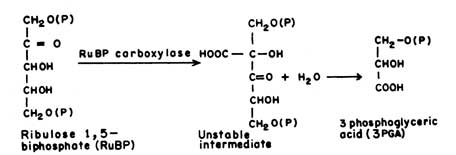
(3-Phospho D-glycerate Carboxylyase EC 4.1.1.39)Ribulose biphosphate carboxylase (RuBP-carboxylase) is the key enzyme which catalyses the fixation of CO2 in plants. It catalyses the combination of RuBP and CO2 resulting in the formation of 3-phosphoglyceric acid.
 |
Apart from the synthetic pathway, portion of 3-phosphoglyceric acid is recycled to form RuBP. The plants which produce C3 acids (3PGA) are termed as C3 plants which include wheat, spinach, bean etc. The fact that RuBP carboxylase constitutes about 50% of soluble leaf protein shows how important this enzyme is to the plant.
RuBP carboxylase also functions as an oxygenase at elevated levels of O2 which seems to be the prime reaction for photorespiration.
Principle
RuBP-carboxylase is assayed radiometrically. The enzyme is made to utilize labeled CO2 as the substrate and the radioactivity in the products is counted as a measure of enzyme activity.
Materials
» HEPES Buffer 25mM,pH 7.8Dissolve 650mg sodium salt of HEPES in water, adjust to pH 7.8 with NaOH and make up to 100mL.
» RuBP 1mM.
Dissolve 4mg tetra sodium salt of RuBP in 10mL water
» MgCl2 1M
Dissolve 952mg in 100mL of water
» Dithiothreitol (DTT), 50mM
Dissolve 771mg in 100mL water
» NaH14CO3 - sp. activity 1m Ci/m mole
10% Acetic Acid (v/v)
» Enzyme Extract
Prepare extraction medium by dissolving 95mg MgCl2 (10mM), 154mg DTT (10mM), 126mg MnCl2 (10mM) and 1.3g sodium salt of HEPES in about 75mL distilled water, adjust to pH 7.8 with NaOH and then make up to 100mL.
Grind 5g leaf sample with a mortar and pestle in about 20mL of the ice cold extraction medium. Centrifuge the homogenate at 20,000g for 10 minutes. Use the supernatant.
Procedure
| 1. |
Prepare a reaction mixture with the following recipes in a tube: 0.10mL HEPES buffer, 0.25mM, pH 7.8. 0.05mL RuBPl; 1mM. 0.05mL MgCl2,100mM. 0.05mLDTT,50mM. 0.20mL enzyme extract. |
| 2. |
Illuminate the reaction mixture at 25°C for 2min. |
| 3. |
Initiate the reaction by adding 1mCi/m mole of NaH14CO3. |
| 4. |
Incubate the assays for 10min. |
| 5. |
Stop the reaction by adding 0.1mL of 10% acetic acid. |
| 6. |
Transfer known aliquots on to Whatman No.3 filter paper discs, dry and count for radioactivity. |
| 7. |
Run a blank similarly without RuBP and instead add 0.05mL of buffer. |
Calculation
The RuBPcase activity is expressed in terms of mmol CO2 or mmol CO2 per kg protein per second. It is also expressed per kg chlorophyll or m.sq. of the leaf area.Notes
1. The enzyme assay can also be conveniently carried out directly in liquid scintillation vials sealed with serum stoppers. The initiation of the reaction with NaH14CO3 may be done by injecting it through the serum stopper. Likewise, acetic acid may be injected to stop the reaction. The vials are unstoppered in the fume hood and the contents taken to dryness preferably in a forced draft oven at 90°C. One mL water is added followed by 12mL of Liquid Scintillation Cocktail [LSC] solution. The vials are capped and the acid stable (3 PGA) radioactivity is determined by Liquid Scintillation Counting.2. The specific radioactivity of the NaH14CO3 stock is determined by adding a 30mL aliquot (15mmoles) to an LSC vial containing 1mL water previously made basic with three drops of ethanolamine, followed by 12mL of LSC solution.
3. Various buffers and reaction mixtures are also successfully used for the assay of RuBP case.
References
1. Kung, S D, Chollet, R and Marsho, T V (1980) In: Methods in Enzymology (Ed Anthony San Pietro) Academic Press New York 69 p 326.2. Procedure Manual Plant Sciences Division, School of Biological Sciences, M K University Madurai (1982).




