Fig. 37.5. Three categories of transcriptional repression (TAF = trans-activation factor; TIC = transcription initiation complex; TRF = trans-repressing factor).
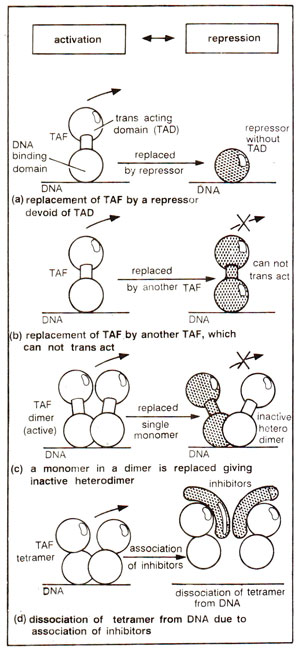
Fig. 37.6. Four ways in which DNA binding of TAF can be inhibited.
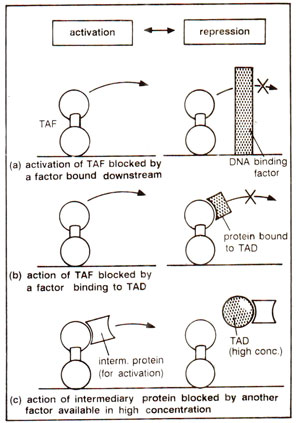
Fig. 37.7. Three ways in which activation (not binding on DNA) by TAF is blocked.
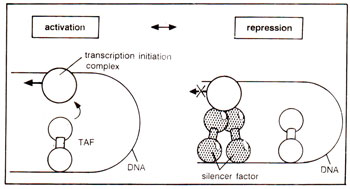
Fig. 37.8. Mehcanism of the action of silencer factors, rendering the transcription initiation complex ineffective.
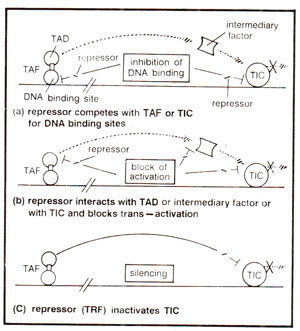
In recent years, many examples of negative control of transcription have also become available in eukaryotes. These examples can be divided into three main classes : (i) inhibition of DNA binding, (ii) blocking of activation and (iii) silencing. These three mechanisms of repression are illustrated in Figure 37.5.

Fig. 37.5. Three categories of transcriptional repression (TAF = trans-activation factor; TIC = transcription initiation complex; TRF = trans-repressing factor).
Inhibition of the binding of trans-activating factors (TAF) to DNA. One of the first examples of inhibition of TAF binding involved
SV40 T antigen binding to its own promoter as in prokaryotes. In many other cases, there is a competition between a negative factor and a positive TAF as shown in the following examples : (i) In sea urchin, sperm specific histone gene
H2B-1 is repressed in embryos, due to interference by a negative factor with a positive CCAAT binding factor, (ii) In humans, fetal γ
-globin gene is repressed in adults due to binding of a factor
NF-E, which inhibits binding of positive factor
CP1 to CCAAT box. (iii) Several genes are repressed, when
GC-box factor competes with
Spl for binding to GC-rich sequences. During such an inhibition, a positive factor for one gene may also function as negative factor for another gene due to similar DNA sequences. This has been shown in several developmental genes in
Drosophila and also for glucocorticoid receptor molecule, (iv) In still other cases, a component (e.g.
Fos) of a heterodimer (e.g.
Jun-Fos) can be replaced to give inactive complex (e.g.
C-Jun-Jun B) which competes with heterodimers, or an inhibitor (e.g.
IkB) may cause a TAF dessociate from DNA and inactivate it (Fig. 37.6). Heat shock protein
(hsp 90) also associates with glucocorticoid receptor and renders it incapable of binding DNA.

Fig. 37.6. Four ways in which DNA binding of TAF can be inhibited.
Blocking of activation. In several examples,
TAF can bind DNA, but its activation domain is not allowed to function (Fig. 37.7). Following alternatives are possible : (i) Negative factor may bind near the binding sites meant for positive factors. For instance, in
c-myc gene, the negative factor
myc-PRF binds next to positive factor
myc-CFl, and for polyoma virus, repressor
PEA2 binds near
PEAl. (ii) Negative factor may bind the positive factor, rendering its trans-activation domain ineffective as done by binding of
GAL-80 to
TAF GAL4 in yeast;
similar is the situation with binding of
AP-1 with
P67SRF needed for activation of
C-fos gene (AP-1 is usually a TAF, but here it works as a repressor). (iii) In signal transduction, sometimes an intermediary protein is involved, which is blocked by another trans-activating domain, which may be present in high concentration (e.g. receptors of several steroid hormones). Ela (an adenovirus protein immortalizing primary cells) also blocks activation of several genes by binding intermediary proteins involved in activation due to TAFs (AP-1, GAL4, VP16), which bind to SV40 enhancers.

Fig. 37.7. Three ways in which activation (not binding on DNA) by TAF is blocked.
Silencing of transcription. After the discovery of enhancer elements involved in enhancing transcription in 1985, elements were discovered in yeast, which mediated repression and therefore were called
silencers. Like an enhancer, a silencer also functions irrespective of its position (upto 2.5kb away) and orientation relative, to the gene, whose expression it controls. The role of silencers can be illustrated using the example of mating type loci in yeast, where a mating type active locus
MAT is associated with two silent loci, one
HMR on the right side and the other
HML on the left side. These silent loci can be 'α'
mating type or
'a' mating type and can replace
MAT to bring about a change in mating type (see later in this section).
The loci
HMR and
HML are repressed by silencer located ~ 1 kb upstream in each case and described as
ER (or
HMR-E)and
EL (or
HML-E)
. HMR-E has been shown to have binding sites for a transcription factor
RAP-1 and another protein
ABF-1 needed at replication origin. Both proteins are needed for repression. Similarly α
2 protein coded by one of the genes at
HML locus binds to
HML-E and represses mating type gene(s). Silencer sequences have also been reported for human
immunodeficiency virus (HIV) and called
negative response element (NRE), and also for chicken lysozyme gene. The silencer factor (a protein) either locks the transcription initiation complex and makes it unavailable for activating factors or it disorganizes the transcription initiation complex (Fig. 37.8).

Fig. 37.8. Mehcanism of the action of silencer factors, rendering the transcription initiation complex ineffective.
It should be realized that repression of gene activity is not mediated by a single mechanism and one or more of the above three mechanisms may be involved in the same case or even additional mechanisms may be involved. It has been shown that the steroid receptor superfamily utilizes all the three mechanisms for repression and is also involved in activation of transcription although of different genes. More information in this connection will become available in coming years.










