Archae
The Archaea are a group of single-celled microorganisms. A single individual or species from this domain is called an archaeon (sometimes spelled "archeon"). Archaea, like bacteria, are prokaryotes. They have no cell nucleus or any other organelles within their cells. In the past they were viewed as an unusual group of bacteria and named archaebacteria but since the Archaea have an independent evolutionary history and show many differences in their biochemistry from other forms of life, they are now classified as a separate domain in the three-domain system. In this system, introduced by Carl Woese, the three main branches of evolutionary descent are the Archaea, Eukaryota and Bacteria. Archaea are further divided into four phyla of which two, the Crenarchaeota and the Euryarchaeota, are most intensively studied. Classifying the Archaea is still difficult, since the vast majority of these organisms have never been studied in the laboratory and have only been detected by analysis of their nucleic acids in samples from the environment.Generally, archaea and bacteria are quite similar in size and shape, although a few archaea have very unusual shapes, such as the flat and square-shaped cells of Haloquadra walsbyi. Despite this visual similarity to bacteria, archaea possess genes and several metabolic pathways that are more closely related to those of eukaryotes: notably the enzymes involved in transcription and translation.
Archaea Fossil range: Paleoarchean - Recent |
|
 |
|
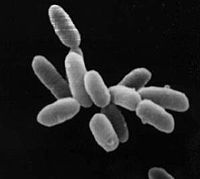
Halobacteria sp. strain NRC-1, each cell about 5 μm in length. |
|
Other aspects of archaean biochemistry are unique, such as their reliance on ether lipids in their cell membranes. The archaea exploit a much greater variety of sources of energy than eukaryotes: ranging from familiar organic compounds such as sugars, to using ammonia, metal ions or even hydrogen gas as nutrients. Salt-tolerant archaea (the Halobacteria) use sunlight as a source of energy, and other species of archaea fix carbon; however, unlike plants and cyanobacteria, no species of archaea is known to do both. Archaea reproduce asexually and divide by binary fission, fragmentation, or budding; in contrast to bacteria and eukaryotes, no species of archaea are known that form spores.
Initially, archaea were seen as extremophiles that lived in harsh environments, such as hot springs and salt lakes, but they have since been found in a broad range of habitats, such as soils, oceans, and marshlands. Archaea are particularly numerous in the oceans, and the archaea in plankton may be one of the most abundant groups of organisms on the planet. These prokaryotes are now recognized as a major part of life on Earth and may play an important role in both the carbon cycle and nitrogen cycle. No clear examples of archaeal pathogens or parasites are known, but they are often mutualists or commensals. One example are the methanogenic archaea that inhabit the gut of humans and ruminants, where they are present in vast numbers and aid in the digestion of food. Archaea have some importance in technology, with methanogens used to produce biogas and as part of sewage treatment, and enzymes from extremophile archaea that can resist high temperatures and organic solvents are exploited in biotechnology.
Contents
Classification
A new domain Early in the 20th century, prokaryotes were regarded as a single group of organisms and classified based on their biochemistry, morphology and metabolism. For example, microbiologists tried to classify microorganisms based on the structures of their cell walls, their shapes, and the substances they consume. However, a new approach was proposed in 1965, using the sequences of the genes in these organisms to work out which prokaryotes are genuinely related to each other. This approach, known as phylogenetics, is the main method used today.Archaea were first classified as a separate group of prokaryotes in 1977 by Carl Woese and George E. Fox in phylogenetic trees based on the sequences of ribosomal RNA (rRNA) genes. These two groups were originally named the Archaebacteria and Eubacteria and treated as kingdoms or subkingdoms, which Woese and Fox termed Urkingdoms. Woese argued that this group of prokaryotes is a fundamentally different sort of life. To emphasize this difference, these two domains were later renamed Archaea and Bacteria. The word archaea comes from the Ancient Greek ἀρχαῖα, meaning "ancient things".
At first, only the methanogens were placed in this new domain, and the archaea were seen as extremophiles that exist only in habitats such as hot springs and salt lakes.
 |
|
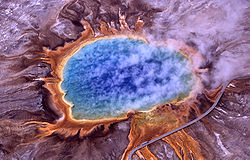
Archaea were first detected in extreme environments, such as volcanic hot springs. |
By the end of the 20th century, microbiologists realized that the archaea are a large and diverse group of organisms that are widely distributed in nature and are common in much less extreme habitats, such as soils and oceans. This new appreciation of the importance and ubiquity of archaea came from using the polymerase chain reaction to detect prokaryotes in samples of water or soil from their nucleic acids alone. This allows the detection and identification of organisms that cannot be cultured in the laboratory, which is often difficult.
Current classification
The classification of archaea, and of prokaryotes in general, is a rapidly moving and contentious field. Current classification systems aim to organize archaea into groups of organisms that share structural features and common ancestors. These classifications rely heavily on the use of the sequence of ribosomal RNA genes to reveal relationships between organisms (molecular phylogenetics). Most of the culturable and well-investigated species of archaea are members of two main phyla, the Euryarchaeota and Crenarchaeota. Other groups have been tentatively created. For example, the peculiar species Nanoarchaeum equitans, which was discovered in 2003, has been given its own phylum, the Nanoarchaeota. A new phylum Korarchaeota has also been proposed, it contains a small group of unusual thermophilic species that shares features of both of the main phyla, but is most closely related to the Crenarchaeota. Other recently detected species of archaea are only distantly related to any of these groups, such as the Archaeal Richmond Mine Acidophilic Nanoorganisms (ARMAN), which were discovered in 2006. |
|
The ARMAN are a new group of archaea recently discovered in acid mine drainage. |
|
The classification of archaea into species is also controversial. In biology, a species is a group of related organisms. A popular definition of a species in animals is a set of organisms that can breed with each other and are reproductively isolated from other groups of organisms (i.e. they cannot breed with other species). However, efforts to classify prokaryotes such as archaea into species are complicated by the fact that they are asexual and show high levels of horizontal gene transfer between lineages. The area is contentious; with, for example, some data suggesting that in archaea such as the genus Ferroplasma, individual cells can be grouped into populations that have highly-similar genomes and rarely transfer genes with more divergent groups of cells. These groups of cells are argued to be analogous to species. On the other hand, studies in Halorubrum found significant genetic exchange between such populations. Such results have led to the argument that classifying these groups of organisms as species would have little practical meaning.
Current knowledge on the diversity of archaea is fragmentary and the total number of archaean species cannot be estimated with any accuracy. Even estimates of the total number of phyla in the archaea range from 18 to 23, of which only 8 phyla have representatives that have been grown in culture and studied directly. Many of these hypothetical groups are known from only a single rRNA sequence, indicating that the vast majority of the diversity among these organisms remains completely unknown. The problem of how to study and classify uncultured microbes is also encountered in the Bacteria.
Origin and evolution
Although probable fossils of prokaryotic cells have been dated to almost 3.5 billion years ago, most prokaryotes do not have distinctive morphologies and the shapes of fossils cannot be used to identify them as Archaea. Instead, chemical fossils, in the form of the unique lipids found in archaea, are more informative because such compounds do not occur in other groups of organisms. Some publications have suggested that the remains of lipids that may be either archaean or eukaryotic were present in shales dating from 2.7 billion years ago, these data have since been questioned. Such lipids have also been detected in rocks dating back to the Precambrian. The oldest known traces of these isoprene lipids come from the Isua district of west Greenland, which include sediments formed 3.8 billion years old and are the oldest on Earth. The origin of Archaea appears very old indeed and the archaeal lineage may be the most ancient that exists on earth.Woese argued that the bacteria, archaea, and eukaryotes each represent a separate line of descent that diverged early on from an ancestral colony of organisms. A few biologists, however, have argued that the Archaea and Eukaryota arose from a group of bacteria. It is possible that the last common ancestor of the bacteria and archaea was a thermophile, which raises the possibility that lower temperatures are "extreme environments" in archaeal terms, and organisms that live in cooler environments appeared later in the history of life on Earth. Since the Archaea and Bacteria are no more related to each other than they are to eukaryotes, this has led to the argument that the term prokaryote has no real evolutionary meaning and should be discarded entirely.
The relationship between archaea and eukaryotes remains an important problem. Aside from the similarities in cell structure and function that are discussed below, many genetic trees group the two together.
 |
|
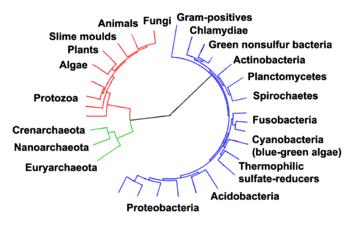
Phylogenetic tree showing the relationship between the archaea and other forms of life. Eukaryotes are colored red, archaea green and bacteria blue. Adapted from Ciccarelli et al. |
Some early analyses even suggested that the relationship between eukaryotes and the archaeal phylum Euryarchaeota is closer than the relationship between the Euryarchaeota and the phylum Crenarchaeota. However, it is now considered more likely that the ancestor of the eukaryotes diverged early from the Archaea. The discovery of archaean-like genes in certain bacteria, such as Thermotoga maritima, makes these relationships difficult to determine, since horizontal gene transfer has occurred. Some have suggested that eukaryotes arose through fusion of an archaean and eubacterium, which became the nucleus and cytoplasm; this accounts for various genetic similarities but runs into difficulties explaining cell structure.
Morphology
Individual archaeans range from 0.1 micrometers (μm) to over 15 μm in diameter, and occur in various shapes, commonly as spheres, rods, spirals or plates. Other morphologies in the Crenarchaeota include irregularly-shaped lobed cells in Sulfolobus, thin needle-like filaments that are less than half a micrometer in diameter in Thermofilum, and almost perfectly rectangular rods in Thermoproteus and Pyrobaculum. There is even a species of flat, square archaea called Haloquadra walsbyi that lives in hypersaline pools. These unusual shapes are probably maintained both by their cell walls and a prokaryotic cytoskeleton. Proteins related to the cytoskeleton components of other organisms exist in the archaea, and filaments are formed within their cells, but in contrast to other organisms, these cellular structures are poorly understood in archaea.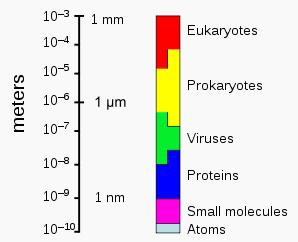 |
|
The sizes of prokaryotes relative to other organisms and biomolecules |
Some species of archaea form aggregates or filaments of cells up to 200 μm in length, and these organisms can be prominent members of the communities of microbes that make up biofilms. An extreme example is Thermococcus coalescens, as aggregates of these cells fuse together in culture, forming single giant cells.
A particularly elaborate form of multicellular colony is produced by archaea in the genus Pyrodictium. Here, the cells produce arrays of long, thin hollow tubes called cannulae that stick out from the cells' surfaces and connect them together into a dense bush-like colony. The function of these cannulae is not known, but they may allow the cells to communicate or exchange nutrients with their neighbors. Colonies can also be produced by an association between different species. For example, in the "string-of-pearls" community that was discovered in 2001 in a German swamp, round whitish colonies of a novel species of archaea in the phylum Euryarchaeota are spaced along thin filaments that can be up to 15 centimetres (5.9 in) long; these filaments are made of a particular species of bacteria.
Cell structure
Archaea are similar to bacteria in their general cell structure, but the composition and organization of some of these structures set the archaea apart. Like bacteria, archaea lack interior membranes so their cells do not contain organelles. They also resemble bacteria in that their cell membrane is usually bounded by a cell wall and they swim by the use of one or more flagella. In overall structure the archaea are most similar to gram-positive bacteria, as most have a single plasma membrane and cell wall, and lack a periplasmic space; the exception to this general rule is the archaean Ignicoccus, which possess a particularly large periplasm that contains membrane-bound vesicles and is enclosed by an outer membrane.Cell membranes
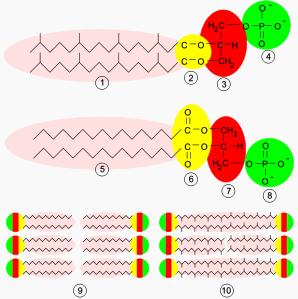
Archaeal membranes are made of molecules that differ strongly from those in other forms of life, which is evidence that archaea are related only distantly to bacteria and eukaryotes. In all organisms cell membranes are made of molecules known as phospholipids. These molecules possess both a polar part that will dissolve in water (the phosphate "head"), and a "greasy" non-polar part that will not dissolve in water (the lipid tail). These dissimilar parts are connected by a glycerol group. In water, phospholipids cluster together, with the polar phosphate heads facing the water and the non-polar lipid tails facing away from the water. This causes them to assemble into layers. The major structure in cell membranes is a double layer of these phospholipids, which is called a lipid bilayer.The phospholipids in the membranes of archaea are unusual in four ways. Firstly, bacteria and eukaryotes have membranes composed mainly of glycerol-ester lipids, whereas archaea have membranes composed of glycerol-ether lipids. The difference between these two types of phospholipid is the type of bond that joins the lipids to the glycerol group; these two types of bonds are shown in yellow in the Figure at the right. In ester lipids this is an ester bond, whereas in ether lipids this is an ether bond. Ether bonds are more chemically-resistant then ester bonds, which might contribute to the ability of some archaea to survive at extremes of temperature and in very acidic or alkaline environments. Bacteria and eukaryotes do contain some ether lipids, but in contrast to archaea these lipids are not a major part of their membranes.
Secondly, archaeal lipids are unique because the stereochemistry of the glycerol group is the reverse of that found in other organisms. The glycerol group can occur in two forms that are mirror images of one another, which may be called the right-handed and left-handed forms; in chemical terms these forms are called enantiomers. Just as a right hand does not fit easily into a left-handed glove, a right-handed glycerol molecule generally cannot be used or made by enzymes adapted for the left-handed form. This suggests that archaea use entirely different enzymes for synthesizing their phospholipids than do bacteria and eukaryotes; since such enzymes developed very early in life's history, this in turn suggests that the archaea split off very early from the other two domains.
Thirdly, the lipid tails of the phospholipids of archaea are chemically different from those in other organisms. Archaeal lipids are based upon the isoprenoid sidechain and are long chains with multiple side-branches and sometimes even cyclopropane or cyclohexane rings. This is in contrast to the fatty acids found in other organisms' membranes, which have straight chains with no branches or rings. Although isoprenoids play an important role in the biochemistry of many organisms, only the archaea use them to make phospholipids. These branched chains may help prevent archaean membranes from becoming leaky at high temperatures.
Finally, in some archaea the phospholipid bilayer is replaced by a single monolayer. In effect, the archaea have fused the tails of two independent phospholipid molecules into a single molecule with two polar heads; this fusion may make their membranes more rigid and better able to resist harsh environments. For example, all the lipids in Ferroplasma are of this type, which is thought to aid this organism's survival in the extraordinarily acidic environments in which it thrives.
Cell wall and flagella
Most archaea possess a cell wall - the exceptions being Thermoplasma and Ferroplasma. In most archaea the wall is assembled from surface-layer proteins, which form an S-layer. An S-layer is made of a rigid array of protein molecules that cover the outside of the cell like chain mail. This layer provides both chemical and physical protection, and can act as a barrier preventing macromolecules from coming into contact with the cell membrane. In contrast to bacteria, most archaea lack peptidoglycan in their cell walls. The exception is pseudopeptidoglycan, which is found in Methanobacteriales, but this polymer is different from the peptidoglycan of bacteria since it lacks D-amino acids and N-acetylmuramic acid.Archaea also have flagella, and these operate in a similar way to bacterial flagella - they are long stalks that are driven by rotatory motors at the base of the flagella. These motors are powered by the proton gradient across the membrane. However, archaeal flagella are notably different in their composition and development. The two types of flagella evolved from different ancestors, the bacterial flagellum evolved from a type III secretion system, while archaeal flagella appear to have evolved from the bacterial type IV pili. In contrast to the bacterial flagellum, which is a hollow stalk and is assembled by subunits moving up the central pore and then adding onto the tip of the flagella, archaeal flagella are synthesized by adding subunits onto their base.
Metabolism
Archaea exhibit a great variety of chemical reactions in their metabolism and use many different sources of energy. These forms of metabolism are classified into nutritional groups, depending on the source of energy and the source of carbon. Some archaea obtain their energy from inorganic compounds such as sulfur or ammonia (they are lithotrophs). These archaea include nitrifiers, methanogens and anaerobic methane oxidisers. In these reactions one compound passes electrons to another (in a redox reaction), releasing energy that is then used to fuel the cell's activities. One compound acts as an electron donor and one as an electron acceptor. A common feature of all these reactions is that the energy released is used to generate adenosine triphosphate (ATP) through chemiosmosis, which is the same basic process that happens in the mitochondrion of animal cells.Other groups of archaea use sunlight as a source of energy (they are phototrophs). However, oxygen-generating photosynthesis does not occur in any of these organisms. Many basic metabolic pathways are shared between all forms of life; for example, archaea use a modified form of glycolysis (the Entner–Doudoroff pathway) and either a complete or partial citric acid cycle. These similarities with other organisms probably reflect both the early evolution of these parts of metabolism in the history of life and their high level of efficiency.
Nutritional types in archaeal metabolism
| Nutritional type | Source of energy | Source of carbon | Examples |
| Phototrophs | Sunlight | Organic compounds | Halobacteria |
| Lithotrophs | Inorganic compounds | Organic compounds or carbon fixation | Ferroglobus, Methanobacteria or Pyrolobus |
| Organotrophs | Organic compounds | Organic compounds or carbon fixation | Pyrococcus, Sulfolobus or Methanosarcinales |
Some Euryarchaeota are methanogens and produce methane gas in anaerobic environments such as swamps. This form of metabolism evolved early, and it is even possible that the first free-living organism was a methanogen. A common reaction in these organisms involves the use of carbon dioxide as an electron acceptor to oxidize hydrogen. Methanogenesis involves a range of coenzymes that are unique to these archaea, such as coenzyme M and methanofuran. Other organic compounds such as alcohols, acetic acid or formic acid are used as alternative electron acceptors by methanogens. These reactions are common in gut-dwelling archaea. Acetic acid is also broken down into methane and carbon dioxide directly, by acetotrophic archaea. These acetotrophs are archaea in the order Methanosarcinales, and are a major part of the communities of microorganisms that produce biogas.
Other archaea use CO2 in the atmosphere as a source of carbon, in a process called carbon fixation (they are autotrophs). In the archaea, this process involves either a highly-modified form of the Calvin cycle, or a recently-discovered metabolic pathway called the 3-hydroxypropionate/4-hydroxybutyrate cycle. The Crenarchaeota also use the reverse Krebs cycle and the Euryarchaeota also use the reductive acetyl-CoA pathway. In these organisms, carbon-fixation is powered by inorganic sources of energy, rather than by capturing sunlight as in plants and cyanobacteria. There are no known archaea that carry out photosynthesis, which is when light is used by photoautotrophs as a source of energy as well as driving the fixation of carbon dioxide. The energy sources used by archaea to fix carbon are extremely diverse, and range from the oxidation of ammonia by the Nitrosopumilales in anammox metabolism to the oxidation of hydrogen sulfide or elemental sulfur by species of Sulfolobus, using either oxygen or metal ions as electron acceptors.
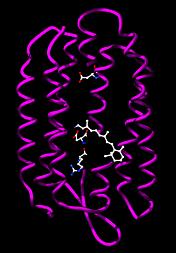
Phototrophic archaea use light to produce chemical energy in the form of ATP. In the Halobacteria, light-activated ion pumps like bacteriorhodopsin and halorhodopsin generate ion gradients by pumping the ions out of the cell across the plasma membrane. The energy stored in these electrochemical gradients is then converted into ATP by ATP synthase. This process is a form of photophosphorylation. The structure and function of these light-driven pumps has been studied in great detail, which has revealed that their ability to move ions across membranes depends on light-driven changes in the structure of a retinol cofactor buried in the center of the protein.
 |
|
Bacteriorhodopsin from Halobacterium salinarum. The retinol cofactor and residues involved in proton transfer are shown as ball-and-stick models. |
|
Genetics
Archaea usually have a single circular chromosome, the size of which may be as great as 5,751,492 base pairs in Methanosarcina acetivorans, the largest archaean genome sequenced to date. At one-tenth of this size is the tiny 490,885 base-pair genome of Nanoarchaeum equitans, which is the smallest archaeal genome known; it is estimated to contain only 537 protein-encoding genes. Smaller independent pieces of DNA, called plasmids, are also found in archaea. Plasmids may be transferred between cells by physical contact, in a process that may be similar to bacterial conjugation.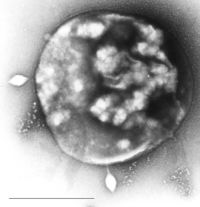 |
|
Sulfolobus infected with the DNA virus STSV1. Bar is 1 micrometer. |
Archaea can be infected by double-stranded DNA viruses that are unrelated to any other form of virus and have a variety of unusual shapes, with some resembling bottles, hooked rods, or teardrops. These viruses have been studied in most detail in the thermophilic archaea, particularly the orders Sulfolobales and Thermoproteales. Defenses against these viruses may involve RNA interference from repetitive DNA sequences within archaean genomes that are related to the genes of the viruses.
Archaea are genetically distinct from bacteria and eukaryotes, with up to 15% of the proteins encoded by any one archaeal genome being unique to the Archaea, although most of these unique genes have no known function. Of the remainder of the genes unique to archaea that have an identified function, most are involved in methanogenesis. The genes that are shared between archaea, bacteria and eukaryotes form a common core of cell function, relating mostly to transcription, translation, and nucleotide metabolism. Other characteristic features of archaean genomes are the organization of genes of related function - such as enzymes catalysing steps in the same metabolic pathway - into novel operons, and large differences in tRNA genes and their aminoacyl tRNA synthetases.
Transcription and translation in archaea are more similar to these processes in eukaryotes than in bacteria, with the archaean RNA polymerase and ribosomes being very close to their equivalents in eukaryotes. Although archaea only have one type of RNA polymerase, its structure and function in transcription seems to be close to that of the eukaryotic RNA polymerase II, with similar assemblies of proteins (the general transcription factors) directing the binding of the RNA polymerase to a gene's promoter. However, other archaean transcription factors are closer to those found in bacteria. Post-transcriptional modification is simpler than in eukaryotes, since most archaean genes lack introns, although there are many introns in their transfer RNA and ribosomal RNA genes, and introns may occur in a few of their protein-encoding genes.
Reproduction
Archaea reproduce asexually by binary or multiple fission, fragmentation, or budding; meiosis does not occur, so if a species of archaea exists in more than one form, these will all have the same genetic material. Cell division is controlled in the archaea in a cell cycle; after the cell's chromosome is replicated and the two daughter chromosomes are separated, the cell divides. The details of the archaeal cell cycle have only been investigated in the genus Sulfolobus, but here it has characters that are similar to both bacterial and eukaryotic systems. In this archaean, the chromosomes are replicated from multiple starting-points (origins of replication) using DNA polymerases that resemble the equivalent eukaryotic enzymes. However, the proteins that direct cell division, such as the protein FtsZ, which forms a contracting ring around the cell, and the components of the septum that is constructed across the center of the cell, are similar to their bacterial equivalents.Spores are made by both bacteria and eukaryotes, but are not formed in any of the known archaea. Some species of Haloarchaea undergo phenotypic switching and grow as several different types of cell, including thick-walled structures that are resistant to osmotic shock and allow the archaea to survive in water at low concentrations of salt, but these are not reproductive structures and may instead help them disperse to new habitats.
Ecology
Habitats
Archaea exist in a broad range of habitats, and are a major part of global ecosystems, and may contribute up to 20% of the total biomass on Earth. Multiple archaeans are extremophiles, and historically this was seen as their ecological niche. Indeed, some archaea survive high temperatures, often above 100 °C, as found in geysers, black smokers, and oil wells. Others are found in very cold habitats and others in highly saline, acidic, or alkaline water. However, other archaea are mesophiles that grow in much milder conditions, in marshland, sewage, the oceans, and soils.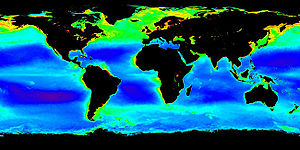 |
|
Image of plankton in the oceans; archaea form a major part of oceanic life. |
Extremophile archaea are members of four main physiological groups. These are the halophiles, thermophiles, alkaliphiles, and acidophiles. These groups are not comprehensive or related to which phylum the organisms belong to, nor are they mutually exclusive, since some archaea belong to several of these groups. Nonetheless, they are a useful starting point for classification.
Halophiles, including the genus Halobacterium, live in extremely saline environments such as salt lakes and start outnumbering their bacterial counterparts at salinities greater than 20-25%. Thermophiles grow best at temperatures above 45°C, in places such as hot springs; hyperthermophilic archaea are defined as those that grow optimally at temperatures greater than 80°C.
The archaeal Methanopyrus kandleri Strain 116 grows at 122 °C, which is the highest recorded temperature at which any organism will grow. Other archaea exist in very acidic or alkaline conditions. For example, one of the most extreme archaean acidophiles is Picrophilus torridus, which grows at pH 0, which is equivalent to thriving in 1.2 Molar sulfuric acid.
This resistance to extreme environments has made archaea the focus of speculation about the possible properties of extraterrestrial life. This has focused on the possibility that microbial life may exist on Mars, and has even led to the suggestion that viable microbes could be transferred between planets in meteorites.
Recently, several studies have shown that archaea exist not only in mesophilic and thermophilic environments but are also present, sometimes in high numbers, at low temperatures as well. For example, archaea are common in cold oceanic environments such as polar seas. Even more significant are the large numbers of archaea found throughout the world's oceans in the plankton community (as part of the picoplankton). Although these archaea can be present in extremely high numbers (up to 40% of the microbial biomass), almost none of these species have been isolated and studied in pure culture. Consequently, our understanding of the role of archaea in the ecology of the oceans is rudimentary, so their full influence on global biogeochemical cycles remains largely unexplored. Some marine Crenarchaeota are capable of nitrification, suggesting these organisms may be important in the oceanic nitrogen cycle, although these oceanic Crenarchaeota may also use other sources of energy. Vast numbers of archaea are also found in the sediments that cover the sea floor, with these organisms making up the majority of living cells at depths over 1 meter into this sediment.
Role in chemical cycling
Archaea are part of the systems on Earth that recycle elements such as carbon, nitrogen and sulfur through the various habitats in ecosystems. Although these activities are vital for the normal function of ecosystems, archaea can also contribute to the changes that humans have made in the environment, and even cause pollution.Archaea carry out many steps in the nitrogen cycle, this includes both dissimilatory reactions that remove nitrogen from ecosystems, such as nitrate-based respiration and denitrification: as well as assimilatory processes that introduce nitrogen, such as nitrate assimilation and nitrogen fixation. The involvement of archaea in ammonia oxidation reactions was recently discovered; these being particularly important in the oceans. The archaea also appear to be crucial for ammonia oxidation in soils, this produces nitrite, which is then oxidized to nitrate by other microbes, and then taken up by plants and other organisms.
In the sulfur cycle, archaea that grow by oxidizing sulfur compounds are important as they release this element from rocks, making it available to other organisms. However, the archaea that do this, such as Sulfolobus, can cause environmental damage since they produce sulfuric acid as a waste product, and the growth of these organisms in abandoned mines can contribute to acid mine drainage.
In the carbon cycle, methanogen archaea are significant as methane producers. The ability of these archaea to remove hydrogen is important in the degradation of organic matter by the populations of microorganisms that act as decomposers in anaerobic ecosystems, such as sediments, marshes and sewage treatment works. However, methane is one of the most abundant greenhouse gases in Earth's atmosphere, constituting 18% of the global total. It is 25 times more potent as a greenhouse gas than carbon dioxide. Methanogens are the primary source of atmospheric methane, and are responsible for most of the world's yearly methane emissions. As a consequence, these archaea contribute to global greenhouse gas emissions and global warming.
Interactions with other organisms
The well-characterized interactions between archaea and other organisms are either mutualism or commensal. As of 2007, no clear examples of archaeal pathogens or parasites are known. However, a relationship has been proposed between the presence of some species of methanogens and infections in the mouth, and Nanoarchaeum equitans may be a parasite of another species of archaea, since it only survives and reproduces within the cells of the Crenarchaeon Ignicoccus hospitalis, and appears to offer no benefit to its host.One well-understood example of mutualism is the interaction between protozoa and methanogenic archaea in the digestive tracts of animals that digest cellulose, such as ruminants and termites. In these anaerobic environments, protozoa break down cellulose from plant material to obtain energy. This process releases hydrogen as a waste product, but high levels of hydrogen will reduce the energy released by this reaction. When methanogens convert the hydrogen to methane, the protozoa benefit as they will gain more energy from breaking down cellulose.
These associations between methanogens and protozoa are taken a step further in several species of anaerobic protozoa, such as Plagiopyla frontata; here the archaea actually reside inside the protozoa and consume the hydrogen produced in their hydrogenosomes. Similar associations with larger organisms are now being found, with the discovery that the marine archaean Cenarchaeum symbiosum lives within (it is an endosymbiont of) the sponge Axinella mexicana.
 |
|
Methanogenic archaea form a symbiosis with termites. |
|
Archaea can also be commensals, benefiting from an association without helping or harming the other organism. For example, the methanogen Methanobrevibacter smithii is by far the most common archaean in the human flora, with this species making up about one in ten of all the prokaryotes in the human gut. As in termites, these methanogens may in fact be mutualists in humans, interacting with other microbes in the gut to aid the digestion of food. Communities of archaea are also associated with a range of other organisms, such as on the surface of corals, and in the region of soil that surrounds plant roots (the rhizosphere).
Significance in technology and industry
Extremophile archaea, particularly those resistant either to heat or to extremes of acidity and alkalinity, are a source of enzymes that function under these harsh conditions. These enzymes have a wide range of uses. For example, thermostable DNA polymerases, such as the Pfu DNA polymerase from Pyrococcus furiosus, have revolutionized molecular biology by allowing the polymerase chain reaction to be used as a simple and rapid technique for cloning DNA. In industry, amylases, galactosidases and pullulanases in other species of Pyrococcus that function at over 100°C allow food processing at high temperatures, such as the production of low lactose milk and whey. Enzymes from these thermophilic archaea also tend to be very stable in organic solvents, allowing their use in environmentally-friendly processes in green chemistry that synthesize organic compounds. The stability of thermophilic enzymes also makes them easier to use in structural biology, consequently the counterparts of bacterial or eukaryotic enzymes from extremophile archaea are often used in structural studies.In contrast to the range of applications of archaean enzymes, the use of the organisms themselves in biotechnology is more restricted. However, methanogenic archaea are a vital part of sewage treatment, since they are part of the community of microorganisms that carry out anaerobic digestion and produce biogas. In mineral processing, Acidophilic archaea display promise for the extraction of metals from ores, including gold, cobalt and copper.
A new class of potentially useful antibiotics has been discovered in archaea. A few of these archaeocins have been characterized, but hundreds more are believed to exist, especially within Haloarchaea and Sulfolobus. These compounds are important since they are different in structure to bacterial antibiotics, so they may have novel modes of action. In addition, they may allow the creation of new selectable markers for use in archaeal molecular biology. The discovery of new archaeocins depends on successful recovery and cultivation of new species of archaea from the environment.
See also
» List of Archaea genera» List of sequenced archeal genomes
References
- "Annual Greenhouse Gas Index (AGGI) Indicates Sharp Rise in Carbon Dioxide and Methane in 2007".- "Trace Gases: Current Observations, Trends, and Budgets". Climate Change 2001. United Nations Environment Programme.
- Agogué H, Maaike B, Dinasquet J, Herndl GJ. (2008). "Major gradients in putatively nitrifying and non-nitrifying Archaea in the deep North Atlantic". Nature 456 (7223): 788–791.
- Albers SV, van de Vossenberg JL, Driessen AJ, Konings WN (September 2000). "Adaptations of the archaeal cell membrane to heat stress". Front. Biosci. 5: D813–20.
- Allers T, Mevarech M (2005). "Archaeal genetics - the third way". Nat. Rev. Genet. 6 (1): 58–73.
- Aravind L, Koonin EV (1999). "DNA-binding proteins and evolution of transcription regulation in the archaea". Nucleic Acids Res. 27 (23): 4658–70.
- archaea. (2008). In Merriam-Webster Online Dictionary.
- Baker, B. J.; Banfield, J. F. (2003), "Microbial communities in acid mine drainage", FEMS Microbiology Ecology 44 (2): 139–152.
- Baker, B.J., Tyson, G.W., Webb, R.I., Flanagan, J., Hugenholtz, P. and Banfield, J.F. (2006). "Lineages of acidophilic Archaea revealed by community genomic analysis. Science". Science 314 (6884): 1933 – 1935.
- Bardy SL, Ng SY, Jarrell KF (February 2003). "Prokaryotic motility structures". Microbiology (Reading, Engl.) 149 (Pt 2): 295–304.
- Barns SM, Delwiche CF, Palmer JD, Pace NR (1996). "Perspectives on archaeal diversity, thermophily and monophyly from environmental rRNA sequences". Proc. Natl. Acad. Sci. U.S.A. 93 (17): 9188–93.
- Barns, Sue and Burggraf, Siegfried. (1997) Crenarchaeota. Version 01 January 1997. in The Tree of Life Web Project
- Based on PDB 1FBB. Data published in Subramaniam S, Henderson R (August 2000). "Molecular mechanism of vectorial proton translocation by bacteriorhodopsin". Nature 406 (6796): 653–7.
- Berg IA, Kockelkorn D, Buckel W, Fuchs G (December 2007). "A 3-hydroxypropionate/4-hydroxybutyrate autotrophic carbon dioxide assimilation pathway in Archaea". Science (journal) 318 (5857): 1782–6.
- Bernander R (1998). "Archaea and the cell cycle". Mol. Microbiol. 29 (4): 955–61.
- Breithaupt H (2001). "The hunt for living gold. The search for organisms in extreme environments yields useful enzymes for industry". EMBO Rep. 2 (11): 968–71.
- Brocks JJ, Logan GA, Buick R, Summons RE (1999). "Archean molecular fossils and the early rise of eukaryotes". Science 285 (5430): 1033–6.
- Bryant DA, Frigaard NU (November 2006). "Prokaryotic photosynthesis and phototrophy illuminated". Trends Microbiol. 14 (11): 488–96.
- Cabello P, Roldán MD, Moreno-Vivián C (November 2004). "Nitrate reduction and the nitrogen cycle in archaea". Microbiology (Reading, Engl.) 150 (Pt 11): 3527–46.
- Cavicchioli R, Curmi P, Saunders N, Thomas T (2003). "Pathogenic archaea: do they exist?". Bioessays 25 (11): 1119–28.
- Chaban B, Ng SY, Jarrell KF (February 2006). "Archaeal habitats—from the extreme to the ordinary". Can. J. Microbiol. 52 (2): 73–116.
- Chappe B, Albrecht P, Michaelis W (July 1982). "Polar Lipids of Archaebacteria in Sediments and Petroleums". Science 217 (4554): 65–66.
- Chelius MK, Triplett EW (April 2001). "The Diversity of Archaea and Bacteria in Association with the Roots of Zea mays L". Microb. Ecol. 41 (3): 252–63.
- Ciaramella M, Napoli A, Rossi M (February 2005). "Another extreme genome: how to live at pH 0". Trends Microbiol. 13 (2): 49–51.
- Ciccarelli FD, Doerks T, von Mering C, Creevey CJ, Snel B, Bork P (2006). "Toward automatic reconstruction of a highly resolved tree of life". Science 311 (5765): 1283–7.
- Coolen MJ, Abbas B, van Bleijswijk J, et al (April 2007). "Putative ammonia-oxidizing Crenarchaeota in suboxic waters of the Black Sea: a basin-wide ecological study using 16S ribosomal and functional genes and membrane lipids". Environ. Microbiol. 9 (4): 1001–16. \
- Damsté JS, Schouten S, Hopmans EC, van Duin AC, Geenevasen JA (October 2002). "Crenarchaeol: the characteristic core glycerol dibiphytanyl glycerol tetraether membrane lipid of cosmopolitan pelagic crenarchaeota". J. Lipid Res. 43 (10): 1641–51.
- Davies PC (1996). "The transfer of viable microorganisms between planets". Ciba Found. Symp. 202: 304–14; discussion 314–7.
- de Queiroz K (2005). "Ernst Mayr and the modern concept of species". Proc. Natl. Acad. Sci. U.S.A. 102 Suppl 1: 6600–7. doi:10.1073/pnas.0502030102.
- De Rosa M, Gambacorta A, Gliozzi A (01 Mar 1986). "Structure, biosynthesis, and physicochemical properties of archaebacterial lipids". Microbiol. Rev. 50 (1): 70–80.
- DeLong EF (1998). "Everything in moderation: archaea as 'non-extremophiles'". Curr. Opin. Genet. Dev. 8 (6): 649–54.
- DeLong EF, Karl DM (September 2005). "Genomic perspectives in microbial oceanography". Nature 437 (7057): 336–42.
- DeLong EF, Pace NR (2001). "Environmental diversity of bacteria and archaea". Syst. Biol. 50 (4): 470–8.
- DiMarco AA, Bobik TA, Wolfe RS (1990). "Unusual coenzymes of methanogenesis". Annu. Rev. Biochem. 59: 355–94.
- Eckburg P, Lepp P, Relman D (2003). "Archaea and their potential role in human disease". Infect Immun 71 (2): 591–6.
- Eckburg PB, Bik EM, Bernstein CN, et al (June 2005). "Diversity of the human intestinal microbial flora". Science 308 (5728): 1635–8.
- Egorova K, Antranikian G (2005). "Industrial relevance of thermophilic Archaea". Curr. Opin. Microbiol. 8 (6): 649–55.
- Elkins JG, Podar M, Graham DE, et al (June 2008). "A korarchaeal genome reveals insights into the evolution of the Archaea". Proc. Natl. Acad. Sci. U.S.A. 105 (23): 8102–7.
- Engelhardt H, Peters J (1998). "Structural research on surface layers: a focus on stability, surface layer homology domains, and surface layer-cell wall interactions". J Struct Biol 124 (2–3): 276–302.
- Eppley JM, Tyson GW, Getz WM, Banfield JF (2007). "Genetic exchange across a species boundary in the archaeal genus ferroplasma". Genetics 177 (1): 407–16.
- Francis CA, Beman JM, Kuypers MM (May 2007). "New processes and players in the nitrogen cycle: the microbial ecology of anaerobic and archaeal ammonia oxidation". ISME J 1 (1): 19–27.
- Francis CA, Beman JM, Kuypers MM (May 2007). "New processes and players in the nitrogen cycle: the microbial ecology of anaerobic and archaeal ammonia oxidation". ISME J 1 (1): 19–27.
- Gaasterland T (1999). "Archaeal genomics". Curr. Opin. Microbiol. 2 (5): 542–7.
- Galagan JE, Nusbaum C, Roy A, et al (April 2002). "The genome of M. acetivorans reveals extensive metabolic and physiological diversity". Genome Res. 12 (4): 532–42.
- Gevers D, Dawyndt P, Vandamme P, et al (2006). "Stepping stones towards a new prokaryotic taxonomy". Philos. Trans. R. Soc. Lond., B, Biol. Sci. 361 (1475): 1911–6.
- Giovannoni SJ, Stingl U. (2005). "Molecular diversity and ecology of microbial plankton". Nature 427 (7057): 343–8.
- Golyshina OV, Pivovarova TA, Karavaiko GI, et al (01 May 2000). "Ferroplasma acidiphilum gen. nov., sp. nov., an acidophilic, autotrophic, ferrous-iron-oxidizing, cell-wall-lacking, mesophilic member of the Ferroplasmaceae fam. nov., comprising a distinct lineage of the Archaea". Int. J. Syst. Evol. Microbiol. 50 Pt 3 (3): 997–1006.
- Gouy M, Li WH (May 1989). "Phylogenetic analysis based on rRNA sequences supports the archaebacterial rather than the eocyte tree". Nature 339 (6220): 145–7.
- Graham DE, Overbeek R, Olsen GJ, Woese CR (2000). "An archaeal genomic signature". Proc. Natl. Acad. Sci. U.S.A. 97 (7): 3304–8.
- Gribaldo S, Brochier-Armanet C (2006). "The origin and evolution of Archaea: a state of the art". Philos. Trans. R. Soc. Lond., B, Biol. Sci. 361 (1470): 1007–22.
- Gupta RS (2000). "The natural evolutionary relationships among prokaryotes". Crit. Rev. Microbiol. 26 (2): 111–31.
- Hahn, Jürgen; Pat Haug (1986). "Traces of Archaebacteria in ancient sediments". System Applied Microbiology 7 (Archaebacteria '85 Proceedings): 178–83.
- Hall-Stoodley L, Costerton JW, Stoodley P (2004). "Bacterial biofilms: from the natural environment to infectious diseases". Nat. Rev. Microbiol. 2 (2): 95–108.
- Hanford MJ, Peeples TL (January 2002). "Archaeal tetraether lipids: unique structures and applications". Appl. Biochem. Biotechnol. 97 (1): 45–62. doi:10.1385/ABAB:97:1:45.
- Hara F, Yamashiro K, Nemoto N, et al (2007). "An actin homolog of the archaeon Thermoplasma acidophilum that retains the ancient characteristics of eukaryotic actin". J. Bacteriol. 189 (5): 2039–45.
- Hixon WG, Searcy DG (1993). "Cytoskeleton in the archaebacterium Thermoplasma acidophilum? Viscosity increase in soluble extracts". BioSystems 29 (2–3): 151–60.
- Horn C, Paulmann B, Kerlen G, Junker N, Huber H (15 Aug 1999). "In vivo observation of cell division of anaerobic hyperthermophiles by using a high-intensity dark-field microscope". J. Bacteriol. 181 (16): 5114–8.
- Howland, John L. (2000). The Surprising Archaea: Discovering Another Domain of Life. Oxford: Oxford University Press. pp. 32.
- Huber H, Hohn MJ, Rachel R, Fuchs T, Wimmer VC, Stetter KO. (2002). "A new phylum of Archaea represented by a nanosized hyperthermophilic symbiont". Nature 417 (6884): 27–8.
- Hugenholtz P (2002). "Exploring prokaryotic diversity in the genomic era". Genome Biol. 3 (2):
- Jahn U, Gallenberger M, Paper W, et al (March 2008). "Nanoarchaeum equitans and Ignicoccus hospitalis: new insights into a unique, intimate association of two archaea". J. Bacteriol. 190 (5): 1743–50.
- Javaux EJ (2006). "Extreme life on Earth—past, present and possibly beyond". Res. Microbiol. 157 (1): 37–48.
- Jenney FE, Adams MW (January 2008). "The impact of extremophiles on structural genomics (and vice versa)". Extremophiles 12 (1): 39–50.
- Kandler, O.; König, H. (1998). "Cell wall polymers in Archaea (Archaebacteria)" (PDF). Cellular and Molecular Life Sciences (CMLS) 54 (4): 305–308.
- Karner MB, DeLong EF, Karl DM (2001). "Archaeal dominance in the mesopelagic zone of the Pacific Ocean". Nature 409 (6819): 507–10.
- Kelman LM, Kelman Z (2004). "Multiple origins of replication in archaea". Trends Microbiol. 12 (9): 399–401.
- Klocke M, Nettmann E, Bergmann I, et al (May 2008). "Characterization of the methanogenic Archaea within two-phase biogas reactor systems operated with plant biomass". Syst. Appl. Microbiol. 31: 190.
- Koch A (1998). "How did bacteria come to be?". Adv Microb Physiol 40: 353–99.
- Koga Y, Morii H (2007). "Biosynthesis of ether-type polar lipids in archaea and evolutionary considerations". Microbiol. Mol. Biol. Rev. 71 (1): 97–120. doi:10.1128/MMBR.00033-06.
- Koga Y, Morii H (November 2005). "Recent advances in structural research on ether lipids from archaea including comparative and physiological aspects". Biosci. Biotechnol. Biochem. 69 (11): 2019–34.
- Konneke M, Bernhard AE, de la Torre JR, Walker CB, Waterbury JB, Stahl DA. (2005). "Isolation of an autotrophic ammonia-oxidizing marine archaeon". Nature 437 (7057): 543–6.
- Könneke M, Bernhard AE, de la Torre JR, Walker CB, Waterbury JB, Stahl DA (September 2005). "Isolation of an autotrophic ammonia-oxidizing marine archaeon". Nature 437 (7058): 543–6.
- Kostrikina NA, Zvyagintseva IS, Duda VI. (1991). "Cytological peculiarities of some extremely halophilic soil archaeobacteria". Arch. Microbiol. 156 (5): 344–49.
- Krieg, Noel (2005). Bergey’s Manual® of Systematic Bacteriology. USA: Springer. pp. 21–6. I
- Kunin V, Goldovsky L, Darzentas N, Ouzounis CA (2005). "The net of life: reconstructing the microbial phylogenetic network". Genome Res. 15 (7): 954–9.
- Kuwabara T, Minaba M, Iwayama Y, et al (November 2005). "Thermococcus coalescens sp. nov., a cell-fusing hyperthermophilic archaeon from Suiyo Seamount". Int. J. Syst. Evol. Microbiol. 55 (Pt 6): 2507–14.
- Lake JA (January 1988). "Origin of the eukaryotic nucleus determined by rate-invariant analysis of rRNA sequences". Nature 331 (6152): 184–6.
- Lake JA. (1988). "Origin of the eukaryotic nucleus determined by rate-invariant analysis of rRNA sequences". Nature 331 (6152): 184–6.
- Lange, M.; Westermann, P.; Ahring, B.K. (2005). "Archaea in protozoa and metazoa". Applied Microbiology and Biotechnology 66 (5): 465–474.
- Lanyi JK (2004). "Bacteriorhodopsin". Annu. Rev. Physiol. 66: 665–88.
- Leininger S, Urich T, Schloter M, et al (August 2006). "Archaea predominate among ammonia-oxidizing prokaryotes in soils". Nature 442 (7104): 806–9.\
- Lepp P, Brinig M, Ouverney C, Palm K, Armitage G, Relman D (2004). "Methanogenic Archaea and human periodontal disease". Proc Natl Acad Sci U S a 101 (16): 6176–81.
- Lipp JS, Morono Y, Inagaki F, Hinrichs KU (July 2008). "Significant contribution of Archaea to extant biomass in marine subsurface sediments". Nature 454: 991.
- López-García P, López-López A, Moreira D, Rodríguez-Valera F (July 2001). "Diversity of free-living prokaryotes from a deep-sea site at the Antarctic Polar Front". FEMS Microbiol. Ecol. 36 (2–3): 193–202.
- Lykke-Andersen J, Aagaard C, Semionenkov M, Garrett RA (September 1997). "Archaeal introns: splicing, intercellular mobility and evolution". Trends Biochem. Sci. 22 (9): 326–31.
- Macalady JL, Vestling MM, Baumler D, Boekelheide N, Kaspar CW, Banfield JF (October 2004). "Tetraether-linked membrane monolayers in Ferroplasma spp: a key to survival in acid". Extremophiles 8 (5): 411–9.
- Madigan MT, Martino JM (2006). Brock Biology of Microorganisms (11th ed.). Pearson. pp. 136. I
- Makarova KS, Grishin NV, Shabalina SA, Wolf YI, Koonin EV (2006). "A putative RNA-interference-based immune system in prokaryotes: computational analysis of the predicted enzymatic machinery, functional analogies with eukaryotic RNAi, and hypothetical mechanisms of action". Biol. Direct 1: 7.
- Mehta MP, Baross JA (December 2006). "Nitrogen fixation at 92 degrees C by a hydrothermal vent archaeon". Science (journal) 314 (5806): 1783–6.
- Mojica FJ, Díez-Villaseñor C, García-Martínez J, Soria E (2005). "Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements". J. Mol. Evol. 60 (2): 174–82.
- Mueller-Cajar O, Badger MR (August 2007). "New roads lead to Rubisco in archaebacteria". Bioessays 29 (8): 722–4.
- Nealson KH (January 1999). "Post-Viking microbiology: new approaches, new data, new insights". Orig Life Evol Biosph 29 (1): 73–93.
- Nelson KE, Clayton RA, Gill SR, et al (1999). "Evidence for lateral gene transfer between Archaea and bacteria from genome sequence of Thermotoga maritima". Nature 399 (6734): 323–9.
- Ng SY, Chaban B, Jarrell KF (2006). "Archaeal flagella, bacterial flagella and type IV pili: a comparison of genes and posttranslational modifications". J. Mol. Microbiol. Biotechnol. 11 (3–5): 167–91.
- Nickell S, Hegerl R, Baumeister W, Rachel R (2003). "Pyrodictium cannulae enter the periplasmic space but do not enter the cytoplasm, as revealed by cryo-electron tomography". J. Struct. Biol. 141 (1): 34–42.
- Norris PR, Burton NP, Foulis NA (2000). "Acidophiles in bioreactor mineral processing". Extremophiles 4 (2): 71–6.
- O'Connor EM, Shand RF (January 2002). "Halocins and sulfolobicins: the emerging story of archaeal protein and peptide antibiotics". J. Ind. Microbiol. Biotechnol. 28 (1): 23–31.
- Onyenwoke RU, Brill JA, Farahi K, Wiegel J (2004). "Sporulation genes in members of the low G+C Gram-type-positive phylogenetic branch ( Firmicutes)". Arch. Microbiol. 182 (2–3): 182–92.
- Papke RT, Zhaxybayeva O, Feil EJ, Sommerfeld K, Muise D, Doolittle WF (2007). "Searching for species in haloarchaea". Proc. Natl. Acad. Sci. U.S.A. 104 (35): 14092–7.
- Pikuta EV, Hoover RB, Tang J (2007). "Microbial extremophiles at the limits of life". Crit. Rev. Microbiol. 33 (3): 183–209.
- Prangishvili D, Forterre P, Garrett RA (2006). "Viruses of the Archaea: a unifying view". Nat. Rev. Microbiol. 4 (11): 837–48.
- Prangishvili D, Garrett RA (2004). "Exceptionally diverse morphotypes and genomes of crenarchaeal hyperthermophilic viruses". Biochem. Soc. Trans. 32 (Pt 2): 204–8.
- Preston, C.M.; Wu, K.Y.; Molinski, T.F.; Delong, E.F. (1996). "A psychrophilic crenarchaeon inhabits a marine sponge: Cenarchaeum symbiosum gen. nov., sp. nov". Proc Natl Acad Sci USA 93 (13): 6241–6.
- Rachel R, Wyschkony I, Riehl S, Huber H (March 2002). "The ultrastructure of Ignicoccus: evidence for a novel outer membrane and for intracellular vesicle budding in an archaeon" (PDF). Archaea 1 (1): 9–18.
- Rappé MS, Giovannoni SJ (2003). "The uncultured microbial majority". Annu. Rev. Microbiol. 57: 369–94.
- Rasmussen B, Fletcher IR, Brocks JJ, Kilburn MR (October 2008). "Reassessing the first appearance of eukaryotes and cyanobacteria". Nature 455 (7216): 1101–4.
- Robertson CE, Harris JK, Spear JR, Pace NR (2005). "Phylogenetic diversity and ecology of environmental Archaea". Curr. Opin. Microbiol. 8 (6): 638–42.
- Romano A, Conway T (1996). "Evolution of carbohydrate metabolic pathways". Res Microbiol 147 (6–7): 448–55.
- Rudolph C, Wanner G, Huber R (May 2001). "Natural communities of novel archaea and bacteria growing in cold sulfurous springs with a string-of-pearls-like morphology". Appl. Environ. Microbiol. 67 (5): 2336–44.
- Samuel BS, Gordon JI (June 2006). "A humanized gnotobiotic mouse model of host-archaeal-bacterial mutualism". Proc. Natl. Acad. Sci. U.S.A. 103 (26): 10011–6.
- Sára M, Sleytr UB (2000). "S-Layer proteins". J. Bacteriol. 182 (4): 859–68.
- Sch⇒fer G, Engelhard M, Müller V (September 1999). "Bioenergetics of the Archaea". Microbiol. Mol. Biol. Rev. 63 (3): 570–620.
- Schimel J (August 2004). "Playing scales in the methane cycle: from microbial ecology to the globe". Proc. Natl. Acad. Sci. U.S.A. 101 (34): 12400–1.
- Schink B (June 1997). "Energetics of syntrophic cooperation in methanogenic degradation". Microbiol. Mol. Biol. Rev. 61 (2): 262–80.
- Schiraldi C, Giuliano M, De Rosa M (2002). "Perspectives on biotechnological applications of archaea" (PDF). Archaea 1 (2): 75–86.
- Schleper C, Holz I, Janekovic D, Murphy J, Zillig W (01 Aug 1995). "A multicopy plasmid of the extremely thermophilic archaeon Sulfolobus effects its transfer to recipients by mating". J. Bacteriol. 177 (15): 4417–26.
- Schmidt TM (2006). "The maturing of microbial ecology" (PDF). Int. Microbiol. 9 (3): 217–23.
- Schopf J (2006). "Fossil evidence of Archaean life" (PDF). Philos Trans R Soc Lond B Biol Sci 361 (1470): 869–85.
- Shand RF; Leyva KJ (2008). "Archaeal Antimicrobials: An Undiscovered Country". in Blum P (ed.). Archaea: New Models for Prokaryotic Biology. Caister Academic Press.
- Simon HM, Dodsworth JA, Goodman RM (October 2000). "Crenarchaeota colonize terrestrial plant roots". Environ. Microbiol. 2 (5): 495–505.
- Sota M; Top EM (2008). "Horizontal Gene Transfer Mediated by Plasmids". Plasmids: Current Research and Future Trends. Caister Academic Press.
- Staley JT (2006). "The bacterial species dilemma and the genomic-phylogenetic species concept". Philos. Trans. R. Soc. Lond., B, Biol. Sci. 361 (1475): 1899–909.
- Synowiecki J, Grzybowska B, Zdziebło A (2006). "Sources, properties and suitability of new thermostable enzymes in food processing". Crit Rev Food Sci Nutr 46 (3): 197–205.
- Takai K, Nakamura K, Toki T, Tsunogai U, Miyazaki M, Miyazaki J, Hirayama H, Nakagawa S, Nunoura T, Horikoshi K (2008). "Cell proliferation at 122°C and isotopically heavy CH4 production by a hyperthermophilic methanogen under high-pressure cultivation". Proc Natl Acad Sci USA 105: 10949–54.
- Teske A, Sørensen KB (January 2008). "Uncultured archaea in deep marine subsurface sediments: have we caught them all?". ISME J 2 (1): 3–18.
- Thauer RK (December 2007). "Microbiology. A fifth pathway of carbon fixation". Science (journal) 318 (5857): 1732–3.
- Theron J, Cloete TE (2000). "Molecular techniques for determining microbial diversity and community structure in natural environments". Crit. Rev. Microbiol. 26 (1): 37–57.
- Thomas NA, Bardy SL, Jarrell KF (2001). "The archaeal flagellum: a different kind of prokaryotic motility structure". FEMS Microbiol. Rev. 25 (2): 147–74.
- Trent JD, Kagawa HK, Yaoi T, Olle E, Zaluzec NJ (1997). "Chaperonin filaments: the archaeal cytoskeleton?". Proc. Natl. Acad. Sci. U.S.A. 94 (10): 5383–8.
- Valentine DL (2007). "Adaptations to energy stress dictate the ecology and evolution of the Archaea". Nat. Rev. Microbiol. 5 (4): 316–23.
- van Hoek AH, van Alen TA, Sprakel VS, et al (01 February 2000). "Multiple acquisition of methanogenic archaeal symbionts by anaerobic ciliates". Mol. Biol. Evol. 17 (2): 251–8.
- Vianna ME, Conrads G, Gomes BP, Horz HP (April 2006). "Identification and quantification of archaea involved in primary endodontic infections". J. Clin. Microbiol. 44 (4): 1274–82.
- Walsby, A.E. (1980). "A square bacterium". Nature 283 (5742): 69–71.
- Wang M, Yafremava LS, Caetano-Anollés D, Mittenthal JE, Caetano-Anollés G (2007). "Reductive evolution of architectural repertoires in proteomes and the birth of the tripartite world". Genome Res. 17 (11): 1572–85.
- Watanabe Y, Yokobori S, Inaba T, et al (January 2002). "Introns in protein-coding genes in Archaea". FEBS Lett. 510 (1-2): 27–30.
- Waters E, et al (2003). "The genome of Nanoarchaeum equitans: insights into early archaeal evolution and derived parasitism". Proc. Natl. Acad. Sci. U.S.A. 100 (22): 12984–8.
- Waters E, Hohn MJ, Ahel I, et al (October 2003). "The genome of Nanoarchaeum equitans: insights into early archaeal evolution and derived parasitism". Proc. Natl. Acad. Sci. U.S.A. 100 (22): 12984–8.
- Wegley, L.; Yu, Y.; Breitbart, M.; Casas, V.; Kline, D.I.; Rohwer, F. (2004). "Coral-associated Archaea" (PDF). Marine Ecology Progress Series 273: 89–96.
- Werner F (September 2007). "Structure and function of archaeal RNA polymerases". Mol. Microbiol. 65 (6): 1395–404.
- Woese C (1998). "The universal ancestor". Proc. Natl. Acad. Sci. U.S.A. 95 (12): 6854–9.
- Woese C, Fox G (1977). "Phylogenetic structure of the prokaryotic domain: the primary kingdoms". Proc Natl Acad Sci USA 74 (11): 5088–90.
- Woese CR (01 March 1994). "There must be a prokaryote somewhere: microbiology's search for itself". Microbiol. Rev. 58 (1): 1–9.
- Woese CR, Gupta R (1981). "Are archaebacteria merely derived 'prokaryotes'?". Nature 289 (5793): 95–6.
- Woese CR, Kandler O, Wheelis ML (1990). "Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya". Proc. Natl. Acad. Sci. U.S.A. 87 (12): 4576–9.
- Xiang X, Chen L, Huang X, Luo Y, She Q, Huang L (2005). "Sulfolobus tengchongensis spindle-shaped virus STSV1: virus-host interactions and genomic features". J. Virol. 79 (14): 8677–86.
- Yoshinari S, Itoh T, Hallam SJ, et al (August 2006). "Archaeal pre-mRNA splicing: a connection to hetero-oligomeric splicing endonuclease". Biochem. Biophys. Res. Commun. 346 (3): 1024–32.
- Yutin N, Makarova KS, Mekhedov SL, Wolf YI, Koonin EV (May 2008). "The deep archaeal roots of eukaryotes". Mol. Biol. Evol. 25: 1619.
- Zillig W (December 1991). "Comparative biochemistry of Archaea and Bacteria". Curr. Opin. Genet. Dev. 1 (4): 544–51.
- Zuckerkandl E, Pauling L (1965). "Molecules as documents of evolutionary history". J. Theor. Biol. 8 (2): 357–66.
Further reading
- Blum P (editor) (2008). Archaea: New Models for Prokaryotic Biology. Caister Academic Press.- Cavicchioli R (2007). Archaea: Molecular and Cellular Biology. American Society for Microbiology.
- Garrett RA, Klenk H (2005). Archaea: Evolution, Physiology and Molecular Biology. WileyBlackwell.
- Howland, John L. (2000). The Surprising Archaea: Discovering Another Domain of Life. Oxford: Oxford University Press.
- Lipps G (2008). "Archaeal Plasmids". Plasmids: Current Research and Future Trends. Caister Academic Press.
- Martinko JM, Madigan MT (2005). Brock Biology of Microorganisms (11th ed.). Englewood Cliffs, N.J: Prentice Hall.