Alteration during replication of nucleic acids
(a) Incorporation of base analogues:
5-bromouracil (5-Bu), 5-chlorouracil (5-Cu) and 5-iodouracil (5-Iu) can replace thymine in DNA but, 2-aminopurine (2-AP) is incorporated in such a small amount that it could not be possible to find out which base it replaces. 2,6-diaminopurine is also highly mutagenic, but its effect has not been studied in detail. Although these base analogues are much less inhibitory for bacterial and phage growth than other purine and pyrimidine analogues, yet they are much more mutagenic. Their mutagenic effect has been attributed, therefore, to pairing mistakes due to their incorporation in DNA.
(
i)
5-Bromouracil (
Bu)
or 5-Bromodeoxy-uridine (
BUdr)
. 5-Bu can pair with adenine just like thymine (Fig. 23.2). Occasionally 5-Bu loses a hydrogen atom in position-1 and pairs with guanine instead of adenine (Fig. 23.3). This pairing mistake may occur at the time of
'incorporation' (Fig. 23.4) or after incorporation at the time of
'replication'.
Hence both kinds of transitions, G-C into A-T and A-T into GC should be inducible. Thus although 5-Bu-induced mutations are mainly due to Bu-incorporation, the frequency of forward mutations is not necessarily proportional to the amount of Bu incorporated, since the ratio of mistakes in incorporation to mistakes in replication may vary. The higher frequency of pairing mistakes are due to higher electronegativity of Br in Bu as compared to methyl group in thymine (methyl uracil). The pyrimidine ring gets poorer in electrons and thus can lose H-atom at position-1 more easily than in thymine, thus giving rise to
enol form. This suggests that mistakes in incorporation may be more frequent, since once incorporated, the electronegativity of Bu is reduced and the mistakes in replication will be less frequent (Fig. 23.5). Thus changes GC →AT should be more frequent.
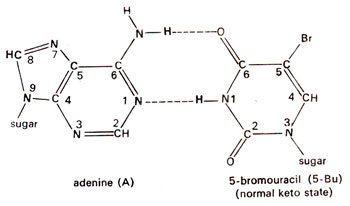
Fig. 23.2. Base pairing between adenine (A) and normal keto state of 5-bromouracil (5-Bu).
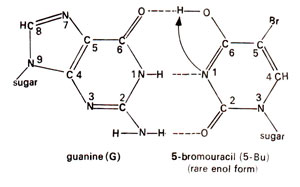
Fig. 23.3. Base pairing between adenine (A) and rare enol form of 5-bromouracil (5-Bu).
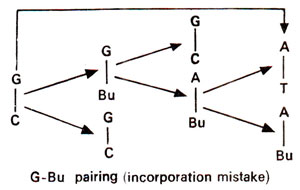
Fig. 23.4. Steps involved in GC →AT base pair transition due to mistake in incorporation of 5-bromouracil (5-Bu).
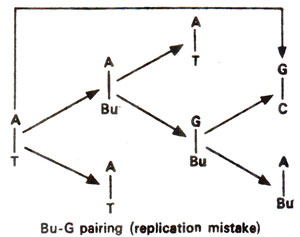
Fig. 23.5. Steps involved in AT → GC base pair transition due to mistake in replication caused by the presence of 5-bromouracil (5-Bu).
Bromodeoxyuridine (BUdR) is more effective mutagen probably because it is more easily converted to deoxyribonucleoside triphosphate. Both Bu and BUdR induce point mutations.
(
ii)
2-Aminopurine (
AP)
. 2-AP can pair with thymine (T) by 2-hydrogen bonds (Fig. 23.6) and with cytosine (C) by a single bond (Fig. 23.7). The pairing involving two bonds is more common, since in the other case, nitrogen (N) in AP and nitrogen (N) in cytosine (C) repel each other and cause their separation. Incorporation of AP at place of guanine (G) to give AP-C base pair will cause mutation in subsequent generations (Fig. 23.8). Similarly a mistake in replication after incorporation of AP, leading to the formation of AP-T base pair may lead to a mutation (Fig. 23.9). AP will thus induce transitions in both directions.
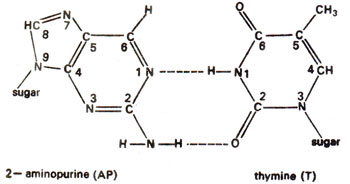
Fig. 23.6. Normal base pairing between 2-aminopurine (2-AP) and thymine (T).
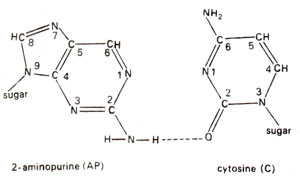
Fig. 23.7. Rare base pairing between 2-aminopurine (2-AP) and cytosine (C).
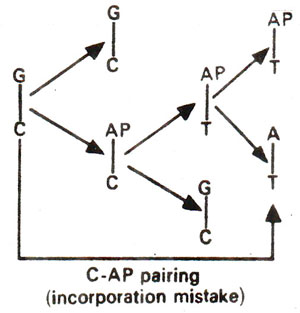
Fig. 23.8. Steps involved in GC →AT base pair transition due to mistake in incorporation of 2-aminopurine (AP).
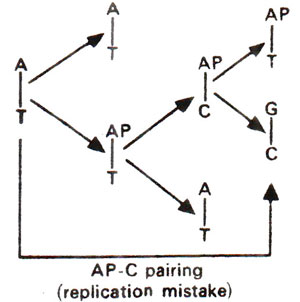
Fig. 23.9. Steps involved in AT →GC base pair transition due to mistake in replication caused by the presence of 2-aminopurine (AP).
(b) Inhibition of nucleic acid precursors. There are several agents which interfere with normal synthesis of nucleic acid precursors. While some interfere with synthesis of purines or pyrimidines, others are more specific and interfere with the synthesis of thymine, adenine, or guanine.
Azaserine is one of the strongest mutagen inhibiting purine synthesis. Since this is also an alkylating agent, its mutagenicity may be due to its alkylating ability rather than its inhibitory action. Similarly
urethane is an inhibitor of pyrimidine synthesis and also a mild alkylating agent. Thymine inhibits the production of chromosome breaks by urethane. It is surprising that adenine also showed mutagenic effect, though normal purine nucleosides have an antimutagenic effect. It is possible that lack of one base either causes breaks or causes pairing mistakes. It is also possible that the inhibitor causes the formation of another base analogue which is the actual mutagen.
Alteration in resting nucleic acid
(a) Nitrous acid: Nitrous acid deaminates, the bases G, C and A with decreasing frequency in DNA and with equal frequencies in RNA. Its mutagenic effect was analysed for tobacco mosaic virus (TMV), bacteria, phages T
2 and T
4, yeast and other organisms. HNO
2 deaminates adenine (A) to hypoxanthine (H) (Fig. 23.10); cytosine (C) to uracil (U) (Fig. 23.11) and guanine (G) to xanthine (X) (Fig. 23.12). H pairs with C (Fig. 23.13); U pairs with A (Fig. 23.14) and X pairs with C (Fig. 23.15). Thus HNO
2 may bring about changes like AT to GC (due to deamination of A to H); GC to AT (due to deamination of C to U) (Figs. 23.16, 23.17), but no change is brought about when guanine is deaminated to xanthine (Fig. 23.18).
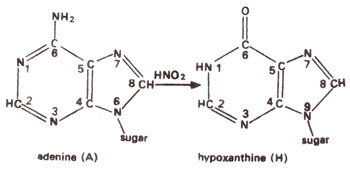
Fig. 23.10. Conversion of adenine (A) into hypoxanthine (H) due to deamination caused by nitrous acid (HNO2).
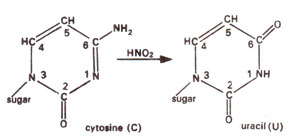
Fig. 23.11. Conversion of cytosine (C) into uracil (U) due to deamination caused by nitrous acid (HNO2).
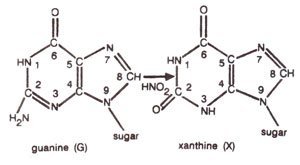
Fig. 23.12. Conversion of guanine (G) into xanthine (X) due to deamination caused by nitrous acid (HNO2).
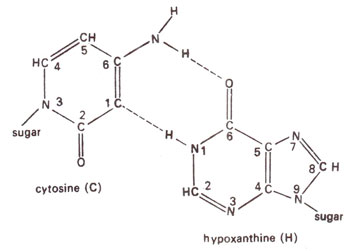
Fig. 23.13. Base pairing between cytosine (c) and hypoxanthine (H).
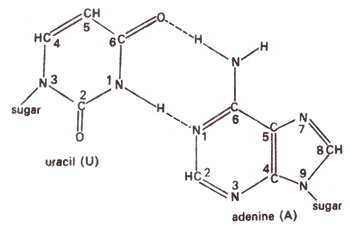
Fig. 23.14. Base pairing between adenine (A) and uracil (U).
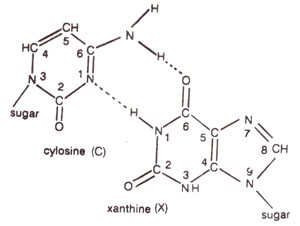
Fig. 23.15. Base pairing between cytosine (C) and xanthine (X)
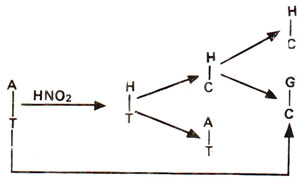
Fig. 23.16. Steps involved in AT → GC base pair transition caused due to deamination of adenine (A) into hypoxanthine (H) and H-C base pairing.
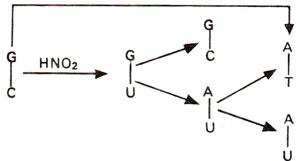
Fig. 23.17. Steps involved in GC→AT base pair transition caused due to deamination of cytosine (C) into uracil (U) followed by A-U base pairing.
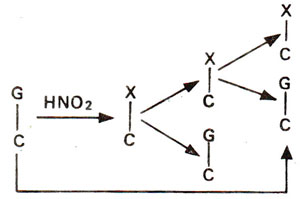
Fig. 23.18. Effect of deamination on guanine (G) to give xanthine (X) resulting into no change in G-C base pair.
(b) Hydroxylamine (HA =
NH2.OH) and hydrazine (HZ =
NH2NH2). In DNA, cytosine is the strongest reacting base when treated with hydroxylamine (HA). Therefore, it is possible that the major mutagenic effect of HA is due to alteration of C. Hydroxylamine probably causes hydroxylation of cytosine at amino group giving rise to hydroxylcytosine, which then subsequently pairs with adenine (Fig. 23.19). This is because the hydroxyl-amino group should be more electronegative than amino group (due to electronegative oxygen) and hence this hydroxylated molecule should be more frequently in the
tautomeric form having a hydrogen atom in place of nitrogen at position-3. This tautomeric form can not pair with guanine (G) but can pair with adenine (A). Thus we should expect that the effect of hydroxylamine (HA) on DNA induces base pair transitions predominantly (Fig. 23.20).
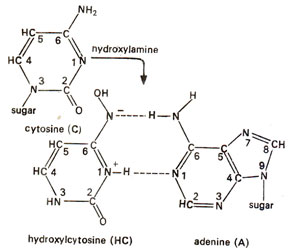
Fig. 23.19. Conversion of cytosine (C) into hydroxylcytosine (HC), which pairs with adenine (A) and not with guanine (G).
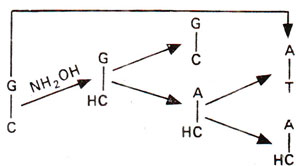
Fig. 23.20. Steps involved in GC→AT base pair transition, due to conversion of cytosine (C) into hydroxylcytosine (HC) followed by A-HC base pairing.
The effect of hydrazine lies in breaking the rings of uracil and cytosine giving rise to
pyrazolone and
3-aminopyrasole respectively. Treatment of RNA by anhydrous hydrazine produces
'ribo-apyrimidinic acid' free of pyrimidines and the treatment of DNA produces the corresponding
'apyrimidinic acid'.
(c) Alkylating agents. Many mutagenic agents carry one, two or more alkyl groups in a reactive form. These are called mono-, bi- or poly functional alkylating agents. Most extensively studied alkylating agents both with tespect to their chemical effects as well as mutagenic effects are diethyl sulphate (DES), dimethyl sulphate (DMS), methyl methane sulphonate (MMS), ethylethane sulphonate (EES) and ethyl methane sulphonate (EMS). They all act as mono-functional agents even when they carry two groups like DES and DMS, since each group alkylates separately. DNA
can be changed by alkylating agents in four different ways.
(i) Alkylation of phosphate groups of nucleic acids. When phosphate group is alkylated (Fig. 23.21), unstable phosphate triester is formed, which hydrolyses to return the alkyl group. If enough alkyl groups remain attached till the time of duplication, the duplication might be inhibited, The attached alkyl group may, but not necessarily, interfere with duplication in such a manner that non-complementary bases may be incorporated.
(ii) Hydrolysis of triester between sugar and phosphate. Hydrolysis of phosphate triester may take place between sugar and phosphate and thus causes the breakage of backbone (Fig. 23.22). This will be lethal or cause larger alterations, but will not cause point mutations.
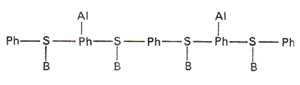
Fig. 23.21. Alkylation (Al) of phosphate (Ph) group in a polynucleotide segment of DNA.
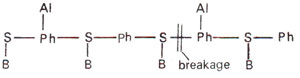
Fig. 23.22. Alkylation (Al) of phosphate (Ph) group leading to breakage of backbone of a polyniicleotide segment of DNA.
(
iii)
Alkylation of bases. Some of the bases are also alkylated (Fig. 23.23). Although 1-methyl adenine, 3-methyl adenine, 1,3-dimethyl adenine and some cytosine derivatives can also be obtained, 7-alkyl guanine is the most common derivative formed, which may pair with thymine instead of pairing with cytosine (Fig. 23.24). The alkylated bases may inhibit DNA duplication or cause base pair mistakes during DNA duplication.
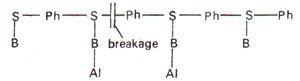
Fig. 23.23. Alkyiation (Al) of bases (B) leading to breakage of backbone of a polynucleotide segment of DNA.
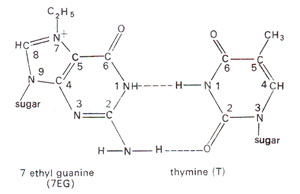
Fig. 23.24. Base pairing between 7-ethylguanine (7-EG) and thymine (T).
(
iv)
Depurination. Alkylation of guanine at 7th position gives rise to quaternary nitrogen which is unstable. This either hydrolyzes away the alky! group or else alkylated purine separates from deoxyribose sugar leaving it depurinated (Fig. 23.25). The gap may then interfere with DNA duplication or may cause incorporation of wrong base (Fig. 23.26). The depurinated DNA is also more labile and may undergo breakage in the backbone soon after. This may induce large alterations or may be lethal but will not cause point mutations.
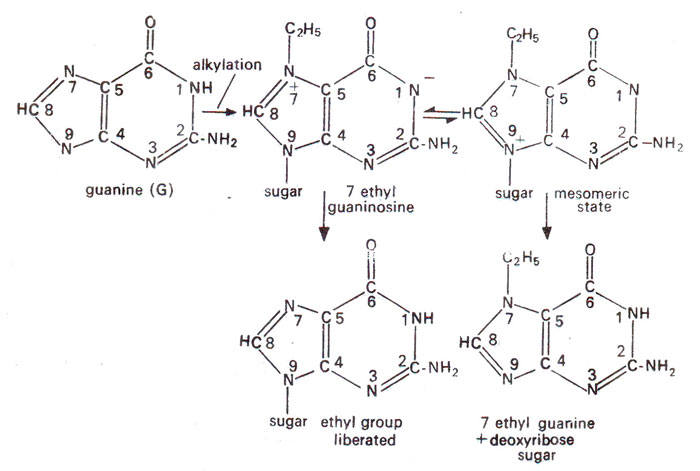
Fig. 23.25. Two different fates of 7-ethylguanine formed due to alkylation : (a) ethyl group liberated or (b) 7-ethylguanine is cleaved off (depurination).
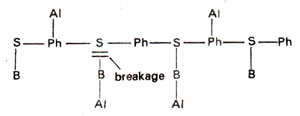
Fig. 23.26. Alkylation of phosphate (Ph) groups and bases (B) leading to breakage of sugar-base linkage.





























