Chromosome Structure
All essential bacterial genes are found in a single, circular, double-stranded DNA chromosome located in the nucleoid region of the cytoplasm. The bacterial chromosome is believed to be attached to the plasma membrane and specifies between 1,000 and 5,000 proteins. It is highly condensed and consists of DNA , RNA, and protein. In addition, there may be one or more plasmids. Plasmids are small circular pieces of extrachromosomal DNA which may encode 20-100 proteins.The genes of eukaryotes are distributed among a number of linear chromosomes that vary in size and number. Eukaryotic chromosomes are condensed by packing the DNA to different degrees (Figure 3-1). Nucleosomes consist of DNA wound twice around an octet of proteins called histones (two each of H2a, H2b, H3, and H4). Approximately 200 base pairs (bp) of the DNA are wound around the spherical bodies formed by the histones, and about 50 bp of DNA connect the nucleosomes. Further compaction may be accomplished by histone H1 binding, which induces the nucleosomes to associate into a ring of six nucleosomes and the rings to associate into a cylinder called a solenoid. Phosphorylation of histone H1 results in the dissociation of the solenoid into an extended nucleosome form. The solenoid is the form in which most of the cell’s DNA exists during interphase. However, further packing can occur by certain proteins binding the solenoid and stimulating it to loop back and forth from a central core of proteins called a scaffold. Dephosphorylation of topoisomerase II and other proteins causes dissociation of the scaffold and results in the decondensation of the chromosomes to the solenoid form. In some eukaryotes, 18 loops of the solenoid form a disklike structure and the chromosome condenses as hundreds of disks stack together. This is the form that is predominant during nuclear division.
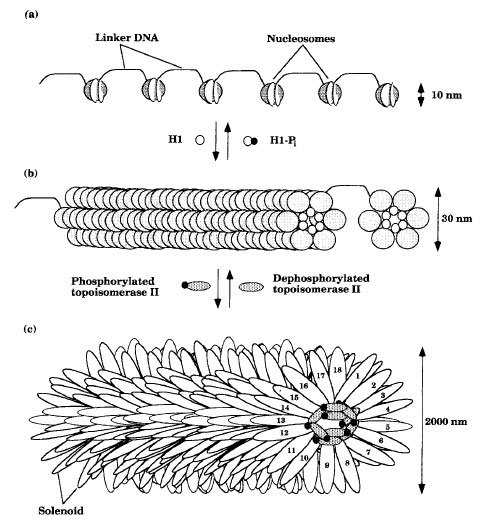 |
| Figure 3-1 Eukaryotic chromosome packaging: (a) extended nucleosome form; (b) solenoid form; and (c) looped solenoid form. |
Heterochromatin is highly condensed DNA that remains in the solenoid form throughout the cell cyle except during DNA replication, when it decondenses. Most of the genes associated with heterochromatin are not expressed because of the DNA ’s condensed state. In contrast, euchromatin is decondensed DNA that exists in the solenoid form or in an extended nucleosome form.
A centromere is a highly constricted region of a mitotic or meiotic chromosome where the spindle fibers attach. Complex sequences of DNA constitute centromeres. If the centromere is in the middle of the chromosome, the chromosome is said to be metacentric. If the centromere is near the tip, it is called telocentric. The short and long arms of the chromosome with respect to the centromere are designated as p and q, respectively. Special staining techniques reveal that each chromosome has a specific pattern of dark and light regions called bands. Homologous chromosomes have the same banding pattern.
Protein complexes associated with the centromeric regions are called kinetochores. Kinetochores bind microtubules of the spindle bundle and function to distribute chromosomes as cells proliferate.
Propagation and maintenance of any piece of DNA requires the presence of one or more origin of replication sites (OriR) and special ends called telomeres. Origin of replication sites are special sequences where DNA replication initiates. Telomeres protect the ends of linear chromosomes from cellular enzymes that degrade nucleic acids from their ends.
Notes
Yeasts have 4 chromosomes; haploid human cells have 23.
Euchromatin in the nucleosome form can be expressed; in the solenoid form it cannot.
Bacterial genomes range from 106 to 3 x 107 bp, whereas a diploid human cell has 5.6 x 109 bp among its 46 chromosomes. However, 90–95% of the human genome is not expressed as protein.




