Genomic Analysis Suggests most Enzymes Evolve from Preexisting Enzymes
The determination of whole genome sequences allowed the identification of all
of the gene families related by primary sequence homology within a specific
organism. Figure 2.1 shows a cluster analysis of the proteins encoded by the
Arabidopsis genome (Thomas Girke, University of California Riverside, personal communication). For example, of the ~27,000 individual proteins in
Arabidopsis,
~80% of proteins are members of homology-related families, whereas only ~20%
represent unique sequences. The distribution shows that approximately half of
the genes are members of groups consisting of >11 members and that nearly one
quarter of proteins belong to groups of >100 members. The larger families include
large numbers of protein kinases and cytochrome
P450s. This clearly illustrates
that new proteins evolved one from another and that divergent evolution is
a primary mechanism for achieving novel functionality.
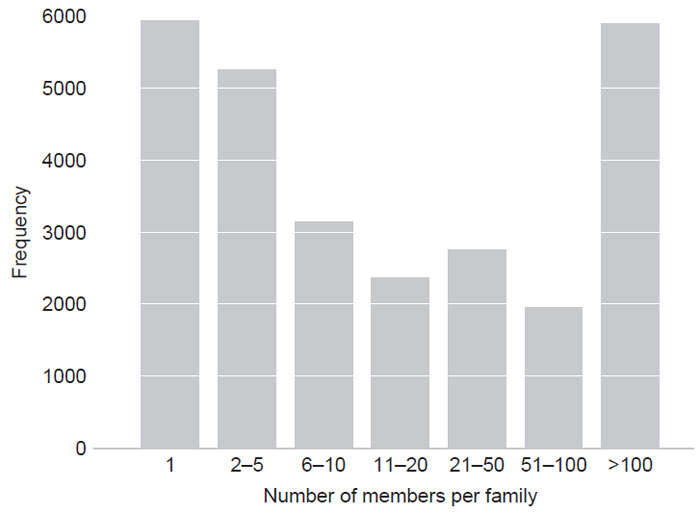 |
| FIGURE 2.1 Frequency distribution of protein families in Arabidopsis. |





