As shown in case of
synthesis of eye pigment in Drosophila, a biosynthetic pathway can be worked out using biochemical mutations, since mutation in a particular gene leads to block in a particular step of biosynthetic pathway. The step which has been blocked can be known either by accumulation of an intermediate product or by recovery of trait due to supply of another intermediate product. This will be illustrated by two additional examples.
Arginine synthesis in Neurospora
In
Neurospora, following three kinds of mutations for arginine synthesis were isolated.
- Mutations, which grow only when arginine is supplied and do not grow with the help of either ornithine or citrulline alone.
- Mutations, which grow only when either citrulline or arginine is supplied, but do not grow when only ornithine is supplied.
- Mutations which can grow only, when either ornithine or citrulline or arginine is supplied.
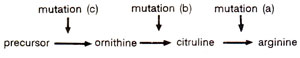
Fig. 22.8. Different steps in the synthesis of arginine and three different mutations blocking three steps.
These mutants indicate that, in the first case : (i) mutants are incapable of utilizing ornithine or citrulline; in the second case, (ii) mutants are incapable of utilizing ornithine; but in the last case, (iii) only the precursor cannot be utilized. The biosynthetic pathway, suggested on the basis of these mutation studies, is shown in Figure 22.8.

Fig. 22.8. Different steps in the synthesis of arginine and three different mutations blocking three steps.
Synthesis of tryptophan in bacteria and Neurospora
Salmonella typhimurium. In
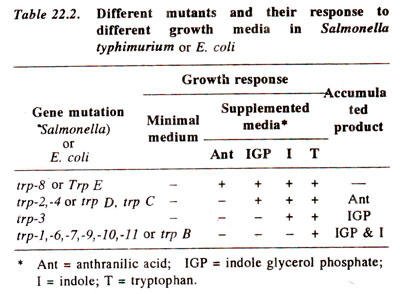
Salmonella typhimurium, four kinds of mutants for tryptophan synthesis could be isolated. These mutations are listed in Table 22.2, and include the following : (i)
trp-8 does not have the ability to synthesize anthranilic acid but the growth can be restored by addition of either of the following compounds—anthranilic acid, indole glycerol phosphate, indole or tryptophan. (ii)
trp-2 and
trp-4 can not grow on either the minimal medium or on the supplemented medium with anthranilic acid, but can grow on addition of either indole glycerol phosphate, or indole or tryptophan. (iii)
trp-3 can grow only on a supplemented medium with either indole or tryptophan, but not on any other medium, (iv)
trp-1, 6, 7, 9, 10, 11, etc. can grow only on a supplemented medium with tryptophan and on no other medium. These different mutants indicate interruption of biochemical pathway at different steps.
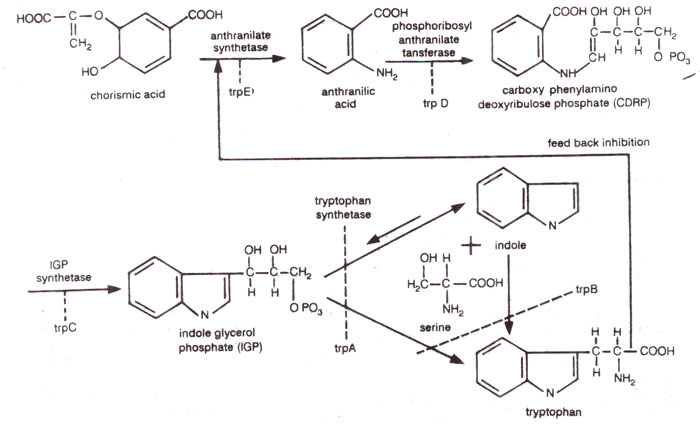
Fig. 22.9. Different steps in the synthesis of amino acid tryptophan, showing the enzymes and genes for each step.
Escherichia coli.In
Escherichia coli also five different kinds of mutations called
trp A, trp B, trp C, trp D and
trp E were identified. These mutations are similar to those described for
Salmonella, and are shown in Figure 22.9 and are summarized in Table 22.2. The biosynthetic pathway shown in Figure 22.9 was traced out using the mutants described above for
Salmonella typhimurium and
Escherichia coli.

Fig. 22.9. Different steps in the synthesis of amino acid tryptophan, showing the enzymes and genes for each step.
Tryptophan synthetase in Neurospora and bacteria. It has been shown that the last two steps of tryptophan metabolism in
Neurospora as well as in bacteria are controlled by the same enzyme
tryptophan synthetase, so that even mutants lacking the ability to utilize indole do not have
tryptophan synthetase enzyme. In
Neurospora, this enzyme consists of only one component. But in
E. coli and
Salmonella, it has two components : (i) 'A' component affecting penultimate step in Figure 22.9, so that
trp A mutants lacking A component can be supplemented by both indole and tryptophan and (ii) 'B' component, which affects last step in Figure 22.9, so that
trpB mutants lacking B component must be supplemented directly with tryptophan. It has also been shown that in mutants for this enzyme, protein is synthesized, but it is unable to perform enzymatic function.
The presence of two components (A and B) of tryptophan synthetase enzyme, synthesized under the control of two different genes suggests that a particular enzyme may consist of more than one polypeptides, each synthesized under the control of a different functional unit called
cistron (see
Fine Structure of Gene-at the Genetic Level (A New Concept of Allelomorphism)). Therefore, it was suggested that the concept of
one gene-one enzyme proposed initially by
Beadle and
Tatum should be modified into
one cistron-one polypeptide hypothesis.









