The embryologists initially studied a developmental process, by destroying, removing, replacing or shifting specific parts of embryo followed by examining the consequences. The same objective can be achieved more conveniently by a geneticist through the study of developmental mutants, characterized by disruption of developmental sequences. Although a number of lethal mutations have been identified in a variety of plants and animals, they only suggest the presence of genes or chromosome segments (some lethal mutations are actually deficiencies for small chromosome segments), which are essential for survival. However, they are not very useful for a study of the developmental process itself. However, there are a number
of other mutations in
Drosophila and some higher plants, which proved to be useful for understanding the genetic control of the development of
different organs.

Fig. 38.3. Ventral view of body segments of Drosophila larva and adult stages (note that each segment found in adult can be identified in the larva).
Developmental mutants in Drosophila
Insect bodies are metameric, consisting of serially repeating units (or body segments), which differentiate into specific structures and patterns according to their position. The generation and diversification of segments during embryo organogenesis depends on two distinct but integrated processes: the subdivision of the embryo into reiterated units, and the specification of their pathway of differentiation. In many insects, segments are generated sequentially as the cells
of the embryo proliferate. By contrast in
Drosophila and other so-called long germ-band insects, the entire body plan is established simultaneously at the blastoderm stage. The adult fly consists of (i) head, (ii) three thoracic segments (prothorax, mesothorax and metathorax, designated as Tl, T2, T3) and (iii) eight abdominals (AB1 to AB8) as shown in Figure 38.3. The T2 carries the wing and the second leg and T3 carries the first leg. These segments can be seen even in the larval stage. A number of
homeotic mutations, disturbing the body plan, have been detected in
Drosophila and used for a detailed study. These homeotic mutations are characterized by the fact, that they convert one part of a body segment into another part of the same segment or into that of a different body segment.

Fig. 38.3. Ventral view of body segments of Drosophila larva and adult stages (note that each segment found in adult can be identified in the larva).
Mutations involving antero-posterior pattern. The antero-posterior pattern is mainly established by the activity of
bicoid (
bed)gene and that of genes of
oskar group. The gene
bcd+ is maternally expressed in the ovary in specialized cells, which form a cluster around the future anterior pole of the developing oocyte. As
bed RNA enters oocyte, it is trapped, apparently by components of cytoskeleton encoded by the genes
swallow (
swa)and
exuperentia (
exu)
, and thus becomes localized at the anterior pole of the oocyte. At about the same time, the products of genes (
nanos = nos gene) of
oskar group get deposited and localized at the posterior end of the egg. Thus the egg gets polarized at the morphological level as well as at the molecular level. Mutants or deficiency for
bed gene produce eggs that develop into embryos with no head or thorax. The removal of products from anterior pole leads to similar results. Similarly, mutants for any
osk group gene develop normal head and thoracic segments but lack entire abdomen. After fertilization,
bed RNA moves backward and
osk RNA moves forward to about the middle of the zygote, each establishing a
gradient. Subsequently, these maternally derived products bind with DNA and thus interact with the zygotic genome.
Gap genes, pair rule genes and homeotic genes. The gap genes and pair rule genes are two classes of genes, which express zygotically before cellularization of blastoderm. The gap genes (the first zygotically active segmentation genes) in their mutant form create gaps in antero-posterior pattern and include
hunchback (
hb)
, krűpal (
kr)and
knirps (
kni)
. Their products have DNA-binding finger domains. The
bed and
osk genes (particularly
nanos)repress transcription of
kr in the anterior and posterior regions respectively, thus restricting its expression to central region. Similarly
bed activates
hb, and
osk activates
kni. Further,
kr and
hb repress each other. This results in the following domains (as revealed using suitable probes) : (i) Expression of
hb in two domains one extending from anterior pole to 50% egg length (EL) and the other extending from posterior pole to 25% of EL; (ii) Expression of
kr is restricted to a broad band in the middle of embryo; (iii) Expression of
kni is restricted to two distinct domains, one anterior and the other posterior to
kr band. These conclusions are based on the study of a number of mutants. These gap genes, in turn, regulate the position specific expression of
pair rule genes and
homeotic genes.
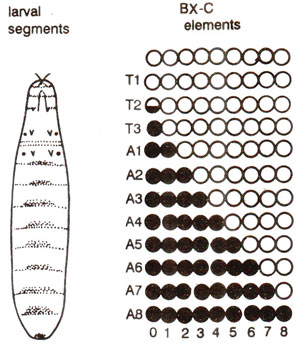
Fig. 38.4. Model of function of bithorax complex; BX-C is divided into nine elements; the active elements; are filled and inactive are open; the half filled cricle represents Ubx+ in the posterior compartment of T2 (note that in each progressively posterior segment, one more element is active).
There are eight pair rule genes (which are expressed periodically), of which six have been characterized and only two (
runt and
hairy = h)appear to have a major function in the generation of the striped patterns of their own expression and that of other genes of this class (hence called pair rule genes). Both genes,
runt and
h also regulate expression of each other. By the last interphase, before cellularization, transcripts of both genes (
runt and
h)are already localized to two complementary series of seven bands that encircle the embryo. The generation of these striped patterns is a key event in the development.
Mutations involving bithorax complex. Mutations in a complex locus,
bithorax complex (
BX-C)in
Drosophila are considered to be the best example for the study of development leading to the production of adult fly. The most important mutations in
BX-C are in
bx+, such as
bx1. For instance, a strong
bx mutation causes the anterior part of T3 to develop as the anterior part of T2. This suggests that
bx+ at some stage becomes active in some cells leading to their differentiation in the anterior part of T3, so that inactivation of
bx+ leads to the development of these cells as anterior of T2. A further extension of the conclusion is that
bx+ does not act in cells that form anterior of T2, since T2 remains unaffected by a
bx mutation. In other words
bx+ remains active in anterior of T3, but turned off in anterior of T2. A deletion in entire
Bx-C is lethal and all segments (T1-T3 and AB1-AB8) develop into those like T2 with its leg and wing, a state which is considered to be the
'developmental ground state'.
Different components of
bx-c at the DNA level are required to direct differentiation of the posterior segments along specialized path of development, so that a recessive mutation affecting a particular segment transforms it to a more anterior segment, which is the ground state for this transformed element. There are also dominant mutations which transform specific segments into structures like those of segments further posterior to them (see Fig. 38.4). Some of the mutants in
Bx-C were described in
Regulation of Gene Expression 3. A Variety of Mechanisms in Eukaryotes.

Fig. 38.4. Model of function of bithorax complex; BX-C is divided into nine elements; the active elements; are filled and inactive are open; the half filled cricle represents Ubx+ in the posterior compartment of T2 (note that in each progressively posterior segment, one more element is active).

Fig. 38.5. The correspondence between antennae and leg structures in Drsophila, based on position specific transformations in homeotic antennae of Antennapedia (mutant Drosophila).
Temperature sensitive mutants. Temperature sensitive (
ts)mutants have also been utilized for the study of development. These mutants may be grown for some time under permissive temperatures and then shifted to restrictive temperatures at specific developmental stages, to study the effect on the phenotype. Pulses of restrictive temperatures have also been used for the analysis of development. For instance, in
Drosophila, the allele,
shibire temperature sensitive 1 (
shits1)was named after a Japanese word meaning paralysis. The mutants with this allele
shi'sl develop normally at 22°C, but when transferred to 29°C they become paralysed and if shifted back to 22°C, the flies recover mobility within minutes. Cultures shifted from 22°C to 29°C at different developmental stages revealed asuccession of different lethal phases (LPs).
A range of phenotypic defects in bristles, hairs and eyes was found to be induced at different times, suggesting the sequence or order in which the developmental stages proceed. Almost all these studies indicated that the temperature sensitive periods occur during the third instar stage of larval development, after which the cells and tissues are committed to
the future course of development, which can not be affected by mutations or by any other manipulation.
Mutations involving antennapedia complex (ANT-C). While
BX-C controls the posterior thoracic and abdominal segments, genes in antennapedia complex (
ANT-C)affect head and thorax. A dominant mutation called
Antennapedia converts part of the antenna into leg structures. This replacement of parts is position-specific, that is, a given internal segment is always replaced by a specific part of the leg (Fig. 38.5). These observations suggest that there is some common overall developmental plan, on which the specific details of leg or antenna development are overlaid.
In these homeotic strains, at the 3' end of
the. gene involved is 180bp (base pairs) long
homeo box, which is associated not only with various genes of
BX-C and
ANT-C complexes (
BX-C and
ANT-C are collectively called homeotic complex or HOM-C), but also with a number of homeotic boxes in frogs, birds, mice and humans. This aspect has been discussed in
Regulation of Gene Expression 3. A Variety of Mechanisms in Eukaryotes.

Fig. 38.5. The correspondence between antennae and leg structures in Drsophila, based on position specific transformations in homeotic antennae of Antennapedia (mutant Drosophila).









