Cell Line Authentication
I. INTRODUCTIONThis article provides a strategy and summarizes steps for the authentication of cell line stocks. It serves as a preface for the following article and for others in this series offering detail on the characterization of cells and cell lines. For those working with serially propagated cells, it is absolutely critical that quality control tests be applied periodically. Rationale and pertinent key references are included here.
Literally hundreds of instances of cross-contamination in cell culture systems have been documented (Nelson-Rees, 1978: Nelson-Rees et al., 1981.; Hukku et al., 1984; MacLeod et al., 1999). Many others have gone unreported. The novice technician or student using cell culture techniques soon is made painfully aware of the potential for bacterial and fungal infection. Generally, however, one must be alerted to the more insidious problems of animal cell cross-contaminations, the presence of mycoplasma, and especially the potential for latent or otherwise inconspicuous viral infection. The financial losses in research and production efforts resulting from the use of contaminated cell lines are certainly equivalent to many millions of dollars. Accordingly, frequent reiteration of the details of cell culture contaminations and of precautionary steps to avoid and detect such problems clearly is warranted.
This article includes a review of quality control steps applied to authenticate cell lines, i.e., to ensure absence of microbial, viral, and cellular contamination, as well as potential tests to verify the identity of human cells. The approach suggested has been developed during the establishment of a national cell repository. Specific rationales for applying the tests indicated are included in this volume and are discussed in more detail elsewhere (Hay et al., 2000).
Most established cell lines have been characterized by the originator and collaborators well beyond the steps essential for quality control. Specific details are provided in subsequent chapters of this series and include, for example, phase-contrast and ultrastructural morphologies; detailed cytogenetic analysis; definition of protooncogene, oncogene, or oncogene product presence, nature, and location; detailed evaluation of intermediate-filament proteins; and demonstration of tissue-specific antigens or production of other specific products. These characterizations obviously increase the value of each line for research and for production work. However, cell resource organizations need not repeat all of these tests before distributing the stock cultures. Decisions must be made to establish the most acceptable authentication steps, consistent with maintaining the lowest possible cost, to provide a highquality cell stock. Authentication can be considered the act of confirming or verifying the identity and critical feature of a specific line, whereas characterization is the definition of the many traits of the cell line, some of which may be unique and also may serve to identify or reauthenticate that line specifically. Essential steps for quality control will vary with the type of cell resource constructed. Minimal descriptive data frequently will be supplemented with a much broader characterization base for each particular cell line.
II. SEED STOCK CONCEPT
Definitions of public repository seed stocks may vary from those used for specific applications such as the production of vaccines or other biologicals. A scheme illustrating the steps involved in developing seed stocks is presented as Fig. 1.
Generally, starter cultures or ampules are obtained from the originator, and progeny are propagated according to the instructions to yield the first "token" freeze. Cultures derived from such token material are then tested for bacterial, fungal, and mycoplasmal contamination. The species of each cell line is verified. These quality control steps are the minimum ones that must be performed before eventual release of a line. If these steps confirm that further efforts are warranted, the material is expanded to produce the seed and distribution stocks. Note that, under ideal conditions, additional major quality control and characterization efforts are applied to cell populations from seed stock ampules. Test results refer to specific numbered stocks. The distribution stock consists of ampules that are distributed on request to investigators. The reference seed stock, however, is retained to generate further distribution stocks as the initial distribution stock becomes depleted. The degree of characterization applied to master cell banks or master working cell banks in production facilities is generally most rigorous. The seed stock here, like the master cell bank, is used as a reservoir to replenish depleted distribution lots over the years. By adherence to this principle, one can avoid problems associated with genetic instability, cell line selection, or transformation.
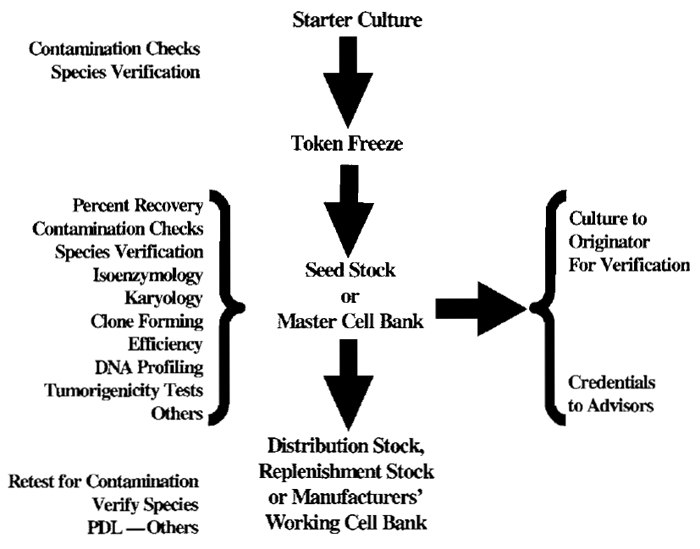 |
| FIGURE 1 Accessioning Scheme. |
III. MICROBIAL CONTAMINATION
Microbial contamination in cell culture systems remains a serious problem. Cryptic contaminants, even of readily isolatable bacteria and fungi, are missed by many laboratories. The American Type Culture Collection (ATCC) still receives cultures, even for the patent depository, that contain yeast, filamentous fungi, and/or mycoplasma contaminants.
A. Bacteria and Fungi
Microscopic examination is not sufficient for the detection of gross contaminations; even some of these cannot be detected readily by simple observations. Therefore, an extensive series of culture tests is also required to provide reasonable assurance that a cell line stock or medium is free of fungi and bacteria. Details are given in the following article.
B. Mycoplasma Infection
Contamination of cell cultures by mycoplasma can be a much more insidious problem than that created by the growth of bacteria or fungi. Although the presence of some mycoplasma species may be apparent because of the degenerative effects induced, other mycoplasma metabolize and proliferate actively in the culture without producing any overt morphological change in the contaminated cell line. Thus, cell culture studies relating to metabolism, surface receptors, virus-host interactions, and so forth are certainly suspect to interpretation, if not negated in interpretation entirely, when conducted with cell lines that harbor mycoplasma. The seriousness of these problems has been documented through published data from testing services and cell culture repositories.
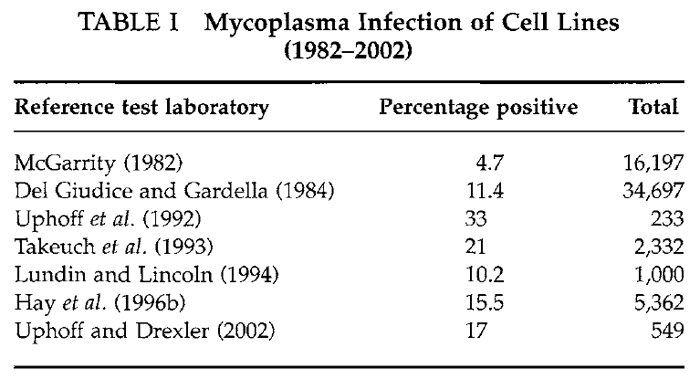
The high incidence of mycoplasma contamination from human operators is supported by the fact that Mycoplasma orale and others of human origin (Mycoplasma hominus, M. salivarium, and M. fermentas) are among those most frequently isolated. In the study of Del Giudice and Gardella (1984) of the 34,697 lines tested, 3955 (11%) were positive; 36% of these isolates were mycoplasmas of human origin. A high incidence of isolation of Mycoplasma hyorhinus was noted that may have resulted from using contaminated sera or by culture-to-culture spread in laboratories working with infected biologicals. After a more recent study, Uphoff et al. (1992) reported that 84 (33%) of 253 cell lines submitted for their developing cell repository in Germany were infected with mycoplasma. Data showing that these are not unusual findings are presented in Table I. Results from seven laboratories published since 1980 indicate clearly that mycoplasma infection is still a very major problem in the cell culture field. Some 5 to 20% of cultures tested were positive. This is the range even today.
Protocols to test for mycoplasma infection are included in the following chapter.
Four general recommendations can be offered to avoid the problem. The implementation of an effective regimen to monitor cell lines for mycoplasma is one critical step. Quarantining all new, untested lines and the use of mechanical pipetting aids are others. Most experts also strongly suggest that the use of antibiotics be eliminated when possible. Antibiotic-free systems permit overgrowth by bacteria and fungi to provide ready indication whenever a lapse in aseptic technique occurs. When a primary tissue is used, e.g., a human tumor sample, antibiotics may be employed initially, but after the primary population has been grown out and cyropreserved, reconstituted cells may be propagated further in antibiotic-free medium.
 |
C. Viruses
Verification of the absence of viruses in cell lines is recognized as a most significant problem. That these may coexist as noncytopathic entities (e.g., the c-type retroviruses) or in a latent form (e.g., papilloma viruses and some herpes viruses) compounds difficulties in detection. Judicious choices are necessary not only to select appropriate methods available for recognizing viruses associated with cell lines, but also to identify the offending species. The nature of the cell resource, its users, and budget available, plus the intended purposes for which the lines will be needed, all affect decisions on testing. More complete detail and protocols are provided in the following article.
IV. CELLULAR CROSS-CONTAMINATION
Wherever cells are grown in culture, serious risk exists for the inadvertent addition and subsequent overgrowth by cells of another individual or species. One most certainly cannot rely on morphologic criteria alone to recognize or identify cell lines. Datadocumenting problems have been collected over the years by groups offering identification services for cell culture laboratories in the United States and elsewhere.
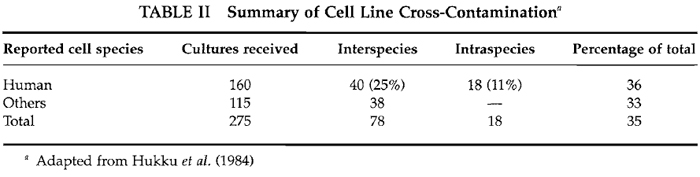
In one study, 466 lines from 62 laboratories were examined. Of these, 75 (16%) were found to be identified incorrectly. A total of 43 lines (9%) were not of the species expected, whereas 32 lines (7%) were either incorrect mixtures of two or more lines or were not the individual line as stated (Nelson-Rees, 1978). Hukku et al. (1984) examined 275 lines over a period of 18 months. Results of their analyses are summarized in Table II. A total of 96 lines (35%) were not as indicated by the donor laboratories. For purported human lines, 36% were not as expected, 25% were a different species, and 11% a different human individual. More recently, Drexler et al. (1994), while developing a resource of cell lines, reported that 10% of those provided by outside investigators contained cells different from those expected, probably due to misidentification or cross-contamination.
 |
To minimize the risk of cellular cross-contamination, culture technicians require a laminar flow hood for ideal operation. These individuals must be instructed periodically to work with only one cell line at any given time, use one reservoir of medium for each line, and avoid introducing pipettes that have been used to dispense or mix cells into any medium reservoir. Technicians must be reminded repeatedly to legibly label each and every cell culture with designations, passages, and dates. Labels of differing colors can be used to more readily distinguish one cell line from another during expansion.
Technicians must also be instructed to allow at least 5min of hood clearance time, with ultraviolet lights and blower on, between cell lines when working on more than one line during a particular period. The inner surfaces of each hood should be swabbed with 70% ethanol between such uses.
The studies outlined earlier illustrate the severity of the problem of cellular cross-contamination and provide a strong rationale for vigilance in careful handling, characterization, and authentication of cell lines.
A. Species Verification
Species of origin can be determined for cell lines by a variety of immunological tests, by isoenzymology, and/or by cytogenetics (Nelson-Rees, 1978; Hukku et al., 1984; Hay et al., 2000). The indirect fluorescent antibody-staining technique is used in many laboratories to verify the species of a cell line (for details, see Hay et al., 1992). Isozyme analyses performed on homogenates of cell lines from over 25 species have demonstrated the utility of these biochemical characteristics for species verification (O'Brien et al., 1977). By determining the mobilities of three isozyme systems - glucose-6-phosphate dehydrogenase, lactic acid dehydrogenase, and nucleoside phosphorylase - using vertical starch gel electrophoresis, the species of origin of cell lines can be identified with a high degree of certainty. Alternatively, a standardized kit employing agarose gels and stabilized reagents may be obtained for this purpose (AuthentiKit, Innovative Chemistry, Inc., Marshfield, MA).
Karyologic techniques have long been used informatively to monitor for interspecies contamination among cell lines. In many instances, the chromosomal constitutions are so dramatically different that even cursory microscopic observations are adequate. In others, for example, in comparisons among cell lines from closely related primates, careful evaluation of banded preparations is required (Nelson-Rees, 1981; Hukku et al., 1984). Cytogenetics has the advantage of detecting even very minor contaminants, on the order of 1% or less in some circumstances. Furthermore, it can provide precise identification of specific lines in cases where marker chromosomes are known or detected (e.g., Drexler et al., 2002). However, cytogenetic analyses are time-consuming and interpretation requires a high degree of skill. The karyotype is constructed by cutting chromosomes from a photomicrograph and arranging them according to arm length, position of centromere, presence of secondary constrictions, and so forth. Automated analytical systems are available but expensive. The "Atlas of Mammalian Chromosomes" (Hsu and Benirschke, 1967-1975) illustrates examples of conventionally stained preparations from over 550 species. Detailed protocols are available elsewhere (Hay et al., 2000).
B. Intraspecies Cross-Contamination
With the dramatic increase in numbers of cell lines being developed, especially from human tissues, the risk of intraspecies cross-contamination rises proportionately. The problem is especially acute in laboratories requiring work with the many different cell lines of human and murine origin available today.
Methods for verifying cell line species employing enzyme mobility studies have been mentioned. Using similar technology, but with different enzyme systems, one can also screen for intraspecies cellular crosscontamination. Cell lines from various individuals of the same species often show different codominant alleles for a given enzyme locus, the products of which are polymorphic and electrophoretically resolvable. In most cases, the phenotype for these allelic enzymes (allozymes) is extremely stable. Consequently, when allozyme phenotypes are determined over a suitable spectrum of loci, they can be used effectively to provide an allozyme genetic signature for each line under study (O'Brien et al., 1977; Hukku et al., 1984).
The application of recombinant DNA technology and cloned DNA probes to identify and quantitate allelic polymorphisms provides additional powerful means for cell line identification. These polymorphisms can be recognized as extremely useful markers, even if they are not expressed through transcription and translation to yield structural enzymatically active proteins.
Hybridization probes to regions of the human genome that are highly variable have been produced for DNA profiling applications, including cell line individualization. Profiles derived from human cell lines can be interpreted best using scanning devices as the patterns are complicated. For protocols and examples, see Jeffreys et al. (1985), Gilbert et al. (1990), and Hay et al. (1996, 2000).
Original procedures included fingerprints using larger (10-20 bp) minisatellite DNA and Southern blotting. With application of the polymerase chain reaction (PCR), loci can be typed in hours rather than days. Smaller microsatellite (2-6bp) loci have been identified as well. Edwards et al. (1992) demonstrated the usefulness of these "short tandem repeat" (STR) loci in recognizing individuals at the DNA level. One significant advantage of STR loci over minisatellite repeats is their small size. The small size of STR loci allows multiplex PCR reactions to be developed in which many loci are examined simultaneously in a single reaction. Commercially available multiplexed STR systems are available for routine screening of new cell lines for authenticity as well as for validating any subsequent distribution of replenishment cell lines. More detail on STR profiling of human DNA cell lines is available elsewhere (Hay et al., 2000). A comprehensive database can be accessed via the ATCC website at http://www.atcc.org/cultures/str.cfm. STR loci are among the most informative polymorphic markers in the genome. The profiling process used at ATCC involves simultaneous amplification of eight STR loci and the amelogenin gene in a multiplex PCR reaction (Promega PowerPlex 1.2 system). This allows for discrimination of fewer than 1 in 108 individuals. The amplicons are separated by electrophoresis and are analyzed using Genotyper 2.0 software from Applied Biosystems. Each peak in the resulting electropherogram represents an allele that is alphanumerically scored and entered into our database.
The classical method for intraspecies cell line individualization involves karyotype analysis after treatment with trypsin and Giemsa stain (Giemsa or G-banding). The banding patterns made apparent by this technique are characteristic for each chromosome pair and permit recognition, by an experienced cytogeneticist, of even comparatively minor inversions, deletions, or translocations. Many lines retain multiple marker chromosomes, readily recognizable by this method, that identify the cells specifically and positively (Chen et al. 1987; Drexler et al. 2002). If readily recognized marker chromosomes are present, contaminations at less than 1% can be recognized with careful scrutiny. This technique even permits discrimination among lines from the same individual that cannot be identified as different by DNA. These lines have similar marker chromosomes, indicating a common source, but each also has markers unique in type and copy number. In contrast, Gilbert et al. (1990) noted no distinct differences among the nine HeLa derivatives examined using the minisattelite probe 33.6 after HaeIII designation. Metabolic differences among the HeLa derivatives (Nelson-Rees et al., 1980) will ultimately be traced to genetic differences among the lines, as reflected by the unique cytogenetic profiles. On this basis, the importance of documenting the precise cytogenetics of cell lines used for production purposes is very clear. Protocols are provided elsewhere in this series.
V. ORIGIN AND FUNCTION
The markers used for verification of the source tissues for cell lines are probably as numerous as the types of metazoan cells. Major methods of demonstration include an analysis of fine structure, immunological tests for cytoskeletal and tissue-specific proteins, and, of course, an extremely broad range of biochemical tests for specific functional traits of tissue cells. Ultrastructural features such as desmosomes or Weibel-Palade bodies identify epithelia and endothelia, respectively. The nature of intermediate-filament proteins, demonstrated using monoclonal antibodies, permits differentiation among epithelial subtypes, mesenchymal, and neurological cells. Tissue- and tumor-specific antigens can be used when reagents are reliable and available. In addition, tissue-specific biochemical reactions or syntheses may be used for absolute identification if these features are retained by the cell line in question. One excellent example is the cell line NCI-H820 (ATCCmHTB 181) isolated from a metastic lesion of a human papillary lung adenocarcinoma. Cells of this line reportedly retain multilamellar bodies suggestive of type 2 pneumocytes and express the three surfactant-associated proteins SP-A (constitutively), SP-B, and SP-C (after dexamethasone stimulation; A. Gazdar, personal communication). For more examples, see other articles of this series and Hay (1992).
VI. CONCLUSIONS
The overall utility of any bank of cultured cell lines depends on the degree of characterization of the holdings that has been performed by the originators, the banking agency, and other individuals within the scientific community. Ready availability at a reasonable cost of the lines and such data, as well as the ability to track distribution of the biologicals, are additional critical considerations. Documenting the verification of species and identity of each cell line, when possible, is considered essential. Freedom from bacterial, fungal, and mycoplasmal infection must be assured. However, from the cell banking perspective, applying all possible characterizations to every seed or master cell stock developed is neither essential nor practical. At ATCC, for example, screens for particular viruses have been applied when a specific program support is available for such testing. Similarly, the definition of ultrastructural features, tumorigenicity, and functional traits, for example, is performed with appropriate external support and adequate rationale. The central responsibility is to produce reference stocks, authenticated and well characterized for multiple purposes, and to return to those preparations over the years for the development of working stocks for distribution or other specific applications. Each replacement distribution stock requires reauthentication prior to distribution to intended users.
References
Chen, T. R. (1988). Re-evaluation of HeLa, HeLaS3 and Hep-2 karyotypes. Cytogenet. Cell Genet. 48, 19-24.
Chen, T. R., Drabkowski, D., Hay, R. J., Macy, M. L., and Peterson, W., Jr. (1987). WiDr is a derivative of another colon adenocarcinoma cell line, HT-29. Cancer Genet. Cytogenet. 27, 125-134.
Del Giudice, R. A., and Gardella, R. S. (1984). Mycoplasma infection of cell culture: Effects, incidence and detection. In "In Vitro Monograph 5: Uses and Standardization of Vertebrate Cell Cultures," pp. 104-115. Tissue Culture Association, Gaithersburg, MD.
Drexler, H. G., Gignac, S. M., Minowada, J. (1994). Hematopoietic cell lines. In "Atlas of Human Tumor Cell Lines" (R. J. Hay, R. Gazdar, and J. G. Park, eds.), pp. 213-250. Academic Press, Orlando.
Drexler H. G., Quentmeier, H., Dirks, W. G., Uphoff, C. C., and MacLeod, R. A. (2002). DNA profiling and cytogenetic analysis of cell line WSU-CLL reveal cross-contamination with cell line REH (pre B-ALL). Leukemia 1868-70.
Edwards, A., Hammond, H. A., Jin, L., Caskey, C. T., and Chakraborty, R. (1992). Genetic variation at five trimeric and tetrameric tandem repeat loci in four human population groups. Genomics 241-253.
Gilbert, D.A, Reid, Y. A., Gail, M. H., Pee, D., White, C., Hay, R. J., and O'Brien, S. J. (1990). Application of DNA fingerprints for cell line individualization. Am. J. Hum. Genet. 47, 499-514.
Hay, R. J. (1992). Cell line preservation and characterization. In "Animal Cell Culture: A Practical Approach" (R. I. Freshney, ed.), 2nd Ed., pp.95-148. IRL Press, New York.
Hay, R. J., Caputo, J., and Macy, M. L. (1992). "ATCC Quality Control Methods for Cell Lines," 2nd Ed. ATCC, Rockville, MD.
Hay, R. J., Reid, Y. A., McClintock. P. R., Chen, T. R., and Macy, M. L. (1996). Cell line banks and their role in cancer research. J. Cell Biochem. Suppl. 27, 1-22.
Hay, R. J., Reid, Y. A., and Miranda, M. (1996b). Advances in methodologies for cell line authentication. In "Culture Collections to Improve the Quality of Life" (R. A. Sampson, J. A. Staplers, D. vander Mei, and A. H. Stouthamer, eds.), pp. 131-137. CVS, Baarm, The Netherlands.
Hay, R. J., Cleland, M. M., Durkin, S., and Reid, Y. A. (2000). Cell line preservation and authentication In "Animal Cell Culture: A Practical Approach" (J. R. W. Masters, ed.), 3rd Ed., pp. 69-103. Oxford University Press, New York.
Hsu, T. C., and Benirschke, K. (1967-1975). "At Atlas of Mammalian Chromosomes." Springer-Verlag, New York.
Hukku, B., Halton, D. M., Mally, M., and Peterson, W. D., Jr. (1984). Cell characterization by use of multiple genetic markers. In"Eukaryotic Cell Cultures" (R. T. Acton and J. D., Lynn, eds.), pp. 13-31. Plenum, New York.
Jeffreys, A. J., Wilson, L., and Thein, S. L. (1985). Hypervariable minisatellite regions in human DNA. Nature 314, 67-73.
Lundin, D. J., and Lincoln, C. K. (1994). Mycoplasmal testing of cell cultures by a combination of direct culture and DNAfluorochrome staining. In Vitro 30A, 111.
MacLeod R. A. E, Dirks, W. G., Kaufmann, M., Matsuo Y., Milch, H., and Drexler, H. G. (1999). Widespread intraspecies crosscontamination of human tumor cell line arising at source. Int. J. Cancer 83:555-563.
McGarrity, G. J. (1982). Detection of mycoplasmal infection of cell cultures. Adv. Cell Culture 2, 9-131.
Nelson-Rees, W. A. (1978). The identification and monitoring of cell line specificity. Progr. Clin. Biol. Res. 20, 25-79.
Nelson-Rees, W. A., Daniels, D. W., and Flandermeyer, R. R. (1981). Cross-contamination of cell lines. Science 212, 446-452.
Nelson-Rees, W. A., Hunter, L., Darlington, G. J., and O'Brien, S. J. (1980). Characteristics of HeLa strains: Permanent vs. variable features. Cytogenet. Cell Genet. 27, 216-231.
O'Brien, S. J., Kleiner, G., Olson, R., and Shannon, J. E. (1977). Enzyme polymorphisms as genetic signatures in human cell culture. Science 195, 1345-1348.
Reid, Y. A., Gilbert, D. A., and O'Brien, S. J. (1990). The use of DNA hypervariable probes for human cell line identification. Am. Type Cult. Collect. Newslett. 10(4), 1-3.
Reid, Y. A., and Lou, X. (1993). The use of PCR-amplified hypervariable regions for the identification and characterization of human cell lines. In Vitro 29A, 120A.
Takeuchi, M., Yoshida, T., Satoh, M., Kuno, H., and Ohno, T. (1993). Survey of mycoplasmal contamination in animal cell lines collected by three cell banks in Japan. Bull. JFCC 9, 13-18.
Uphoff, C. C., Brauer, S., Grunicke, D., Gignae, S. M., MacLeod, R. A. E, Quentmeier, H., Steube, K., Tummler, M., Voges, M., Wagner, B., and Drexler, H. G. (1992). Sensitivity and specificity of the different mycoplasma detection assays. Leukemia 6, 335-341.
Uphoff, C. C., and Drexler, H. G. (2002). Detection of mycoplasma in leukemia-lymphoma cell lines using polymerase chain reaction. Leukemia 16, 289-293.




