RNA Splicing in Metazoans
The central dogma of molecular biology that the information flowfromDNAtoRNAto protein involves colinearity of the sequences of the monomer units is somewhat violated in metazoans because of the presence of interrupted or fragmented genes (Fig. 8). Thus, while the polypeptide sequence is colinear with the codons of the coding sequence in the mRNA, the RNA itself is not collinear with the gene from which it is transcribed. In other words, the gene contains additional intervening sequences called introns, which are transcribed but whose RNA sequence is subsequently removed from the final mRNA containing the coding sequence. The primary gene transcripts of nuclear genomes, called heterogeneous nuclear RNA (hnRNAs), are present in a form of protein-bound particles (ribonucleoprotein particles, or hnRNP). RNA splicing is the process of excising introns from hnRNAs, and contiguous exons are then joined to form mature mRNAs, which are subsequently translocated to cytoplasm and are used as templates for translation (Fig. 8). The cleavage and rejoining occur at specific junctions between exons and introns, so that there are no errors in mature mRNA. First, two adjacent exons are aligned, while the intervening intron is extruded, forming a loop (“lariat”) structure. Then the upstream exon is cleaved and joined to the downstream exon via a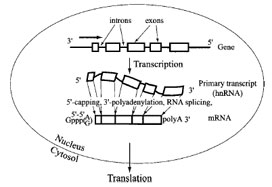 |
| FIGURE 8 A schematic representation of RNA splicing. The coding sequence in metazoan genomes is usually present in segments (exons; indicated by boxes) interspersed between noncoding introns. After synthesis of the primary RNA transcript (called heterogeneous nuclear RNA or hnRNA), the intron sequences are removed by precise cleavage and rejoining is mediated by the spliceosome complex, so that the resulting mature mRNA contains a correctly juxtaposed coding sequence for the polypeptide. The mRNA is also “capped” by 5´-5´ linkage with GMP, and a tail of poly(A) is added at the 3´ terminus to increase the stability of mRNA and to enhance its efficiency in directing protein synthesis when the mRNA is transported from the nucleus to the cytoplasm. |
Termination of eukaryotic transcription is coupled with processing. The mature rRNA is obtained by cleavage of a larger primary transcript synthesized by Pol I. Termination of Pol II transcription occurs at a repeat sequence of U, as in the case of E. coli RNA polymerase, but without the presence of a hairpin structure. More importantly, the 3´ termini of mRNAs are generated by cleavage of primary precursor transcripts followed by addition of a tail of poly(A), a homopolymer of up to several hundred AMP residues synthesized by poly(A) polymerase in a template-independent reaction.




