The lac operon in E. coli
The
lac operator as shown in Figure 35.3, consists of 26 base pairs and is a part of the first structural gene
lac Z (meant for
p galactosidase). There are three structural genes
in lac operon which includes the following : (i) gene
lac Z (3063 bp) codes for the enzyme β
galactosidase, which is active as a tetramer and breaks lactose into glucose and galactose to be utilized in the cell; (ii) gene
lac Y (800 bp) codes for β
galactose permease, which is
a membrane bound protein and helps in transport of metabolites and (iii) gene
lac A (800 bp) which
codes for β
galactose transacetylase, an enzyme that transfers an acetyl group from acetyl-CoA to β galactosides. Any mutations in
lac Z and
lac Y genes give
lac- genotypes, which can not utilize lactose. While
lac Z- mutants abolish enzyme activity,
lac Y- mutants can not take up lactose from
the medium. No defect is observed in
lac A- mutants, which is a puzzle. It is possible that acetylation gives an advantage for utilizing certain non-metabolizable analogues of β-galactoside, because it results in detoxification and excretion.
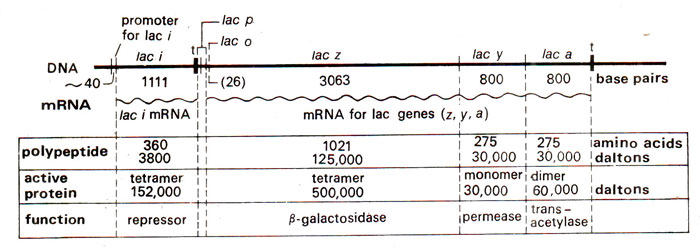

Fig. 35.3. The lactose operon of E. coli and its regulatory gene (redrawn from Lewin's Genes IV).
In
'lac' operon of
E. coli, Tepressor is synthesized through activity of a gene
lac I, known as regulator gene. The repressor then attaches on operator gene and checks synthesis of mRNA (Fig. 35.4). If lactose is added, the repressor is rendered inactive, so that it cannot attach on operator gene, and the synthesis of mRNA takes place (Fig. 35.5). The activity of the regulator gene is controlled by a promoter gene on its left side. It has been shown that mutations in this promoter may increase the quantity of mRNA for repressor. This promoter is different from the promoter for lac operon which is described later in this section.
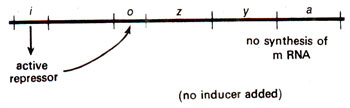
Fig. 35.4. A 'lac' operon showing the action of repressor.
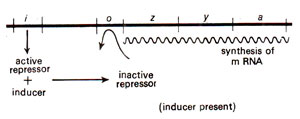
Fig. 35.5. A 'lac' operon showing induction by inducer.
Lactose is not the real inducer of lac operon. Since in the repressed state of
lac operon,
E. coli cells will not synthesize β galactoside permease, one can question how lactose enters the cell. It is believed that a minimal amount
(basal level) of enzyme is always present in the cell to start the metabolic pathway. It has been shown that lactose is not the real inducer of
lac operon, but rather it is an anti-inducer, since it binds to repressor to increase its affinity for operator. On the other hand, the bound protein in inactive repressor was allolactose. It has also been shown that while β galactosidase breaks lactose into glucose and galactose, a side reaction changes galactose into allolactose and galactobiose. Therefore, in the regulation of
lac operon, lactose is taken up and a part of it, through galactose, is converted into allolactose, which then induces the operon, and out-weighs the anti-inducing effect of lactose.
Constitutive mutants. It has been found that a mutation in regulator gene
lac I results in the absence of active repressor and consequently in uncontrolled synthesis of enzyme. Such strains which continuously synthesize enzyme independent of need are known as
constitutive strains or
constitutive mutants. These constitutive strains can also be obtained by mutations in the operator gene. A mutation in operator gene does not allow repressor to act because the repressor functions by acting on operator site. Therefore, there are two types of constitutive strains : (i)
regulator constitutive and
(ii) operator constitutive (Fig. 35.6).
The above two types of mutations, though have the same net result, but differ in their location as well as in their dominant-recessive relationship with respect to the wild type. This relationship has been worked but by using the partial diploid cells (merozygotes). Partial diploid cells are those which have one complete chromosome and a part of another chromosome covering the desired region. When heterozygotes for regulator constitutive were derived, no enzyme synthesis was observed suggesting that wild
lac I is dominant over mutant
lac Ic (Fig, 35.7). On the other hand partial diploid cells heterozygous for mutation at operator site exhibited constitutive synthesis of enzyme, indicating that normal operator gene
'O' is recessive to mutant gene
'Oc' (Fig. 35.8).
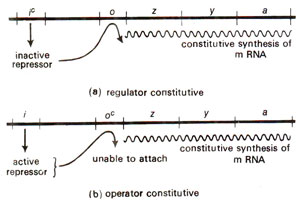
Fig. 35.6. Constitutive strains resulting due to mutations in regulator gene and operator gene (redrawn from Watson : Molecular Biology of the Gene)
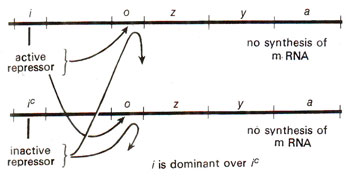
Fig. 35.7. Check on enzyme synthesis in a partial diploid, heterozygous for regulator gene (I/Ic) (redrawn from Watson : Molecular Biology of the Gene).
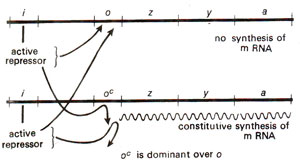
Fig. 35.8. Constitutive enzyme synthesis in a partial diploid, heterozygous for operator gene (O/Oc)(redrawn from Watson : Molecular Biology of the Gene).
There is yet another class of regulator constitutive mutants, which are dominant (
lac Id)
, so that in merozygotes (
lac I/lac Id)
, transcription continues. This
lac ld mutant gives repressor subunit defective at N-terminal region. This defective subunit takes part in formation of a tetramer, which is partly derived from
lac I and partly from
lac Id, so that the tetramer (the form in which repressor binds) is unable to bind on the operator.
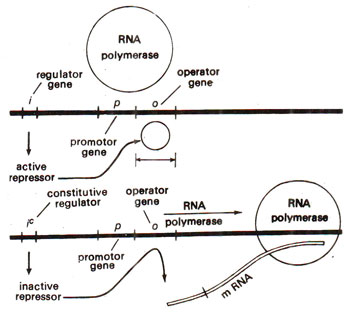
Fig. 35.9. Relationship between promoter gene, repressor and RNA polymerase (redrawn from Watson : Molecular Biology of the Gene).
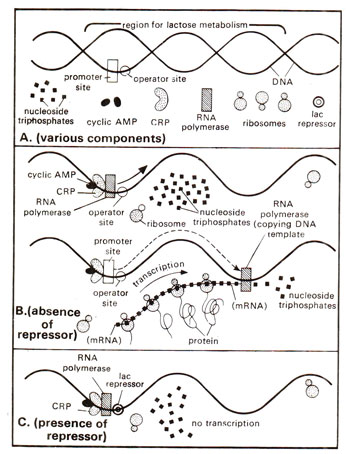
Fig. 35.10. Mechanism involved in the positive control system for the regulation of gene activity in E. coli lac operon. Note that only in the absence of the repressor, RNA polymerase enzyme can travel and transcribe lac operon as shown in B. The repressor, when present on operator site is an obstacle in the path of RNA polymerase.
Promoter gene. While operator gene is the site of action of repressor, the actual site of start of transcription is different and is known as
promoter gene, which lies to the left of operator gene. It is also believed, therefore, that perhaps RNA polymerase binds to and moves from promoter site. The repressor, when attached to operator, prevents the movement of RNA polymerase (Fig. 35.9). The promoter gene in
lac operon was inferred from mutations at this locus, which were found to affect the maximum rate of transcription.

Fig. 35.9. Relationship between promoter gene, repressor and RNA polymerase (redrawn from Watson : Molecular Biology of the Gene).
Thus, there are two proteins which are known
to be involved in the regulation of lactose operon : (i)
lac repressor and
(ii) catabolite gene activator (cga) protein, also known as
CAP (
cyclic AMP protein)
, or
CRP (
cyclic AMP receptor protein)
. While repressor binds to operator site, cga protein binds to cga site. Moreover, repressor exercises a control by checking RNA polymerase to travel through operator, but cga protein exercises a positive control. Only when cyclic AMP molecule activates cga protein, the latter will allow RNA polymerase to bind (Fig. 35.10). A protein such as CAP, which interacts with DNA and RNA polymerase to assist in transcription initiation, is called
apoinducer.

Fig. 35.10. Mechanism involved in the positive control system for the regulation of gene activity in E. coli lac operon. Note that only in the absence of the repressor, RNA polymerase enzyme can travel and transcribe lac operon as shown in B. The repressor, when present on operator site is an obstacle in the path of RNA polymerase.
It has been shown that cyclic AMP is essential for activation of cga protein, since when cyclic AMP was removed or was absent due to mutation, transcription was reduced. In mutants lacking cyclic AMP, transcription was restored by adding cyclic AMP exogenously. It has also been shown that this cyclic-AMP—cga protein system is influenced by the level of glucose concentration. It is, therefore, sometimes described as glucose sensitive system which is common to several glucose sensitive operons in
E. coli, e.g. operons for degradation of lactose, galactose, arabinose, maltose, etc. In all cases glucose is one of the catabolic products of biosynthetic pathway and if accumulates, it is injurious. Therefore, whenever the concentration of glucose increases beyond a certain level, it leads to reduction of transcription. This is achieved by reducing the level of cyclic AMP by glucose, either through checking the synthesis of
adenocyclase, an enzyme needed for synthesis of cyclic AMP or by removing cyclic AMP, by making a complex with it.
Therefore, transcription in
lac operon requires that (i)
cyclic AMP activates
cga protein-a positive control and that (ii)
lac repressor is inactivated by
inducer-a negative control.
Operator and promoter are cis-dominant. It should be realized that in the lac operon, operator can control only
lac genes that are adjacent to it. If a second
lac operon is introduced as done in partial diploids or merozygotes, none of the operators is influenced by the other operator. For instance, if in a partial diploid, one operon has normal operator (
O)and the other has a mutant operator (O
c), the operon with
Oc will be expressed, but that with
O will not be expressed. This could be tested by the using patial diploids which not only differ in operator, but also differ in structural genes.
For instance, in a partial diploid with genotype
OcZ+Y-/OZ-Y+, only β galactosidase will be synthesized, but not the β galactoside permease, although
Y+ is present along with
Oc, but on two differen chromosomes. This phenomenon of lack of dominance extended to other chromosome (trans configuration) is called
cis-dominance. This also suggests that operator does not act through a gene product like regulator gene which acts through repressor, that can diffuse and influence the other allele also. The phenomenon of cis-dominance will be expressed by all those sites, which are physically contiguous with the sequences of controls, and function by being recognized rather than by giving a gene product. It is for this reason that
promoter (described earlier in this section) and
terminator (described in previous section) are also
cis-dominant or
cis-acting, while
regulator is trans-acting.
Uninducible mutants (promoter and regulator genes). Uninducible mutants in the
lac operon are those which can not be induced into synthesis of enzyme even on lactose addition. These can be of two types, cis-acting and trans-acting, (i)
Cis-acting mutations may be in the promoter site (see later), since they act by preventing RNA polymerase from binding to the promoter, so that the operator can not function due to non-availability of RNA polymerase. (ii)
Trans-acting uninducible mutants lie in the regulator gene, since they act by changing the repressor protein so that the altered repressor can not bind the inducer (lactose), and will always be 'locked in' to the active form for attachment to the operator. These mutations are designated as
'lacIs' and are dominant over wild type. This happens because mutant repressor binds to all wild type operators in the cell to prevent their transcription.
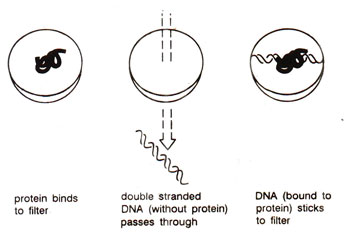
Fig. 35.11. Identification of DNA sequence to which a specific protein is bound, by nitrocellulose filter binding assay.
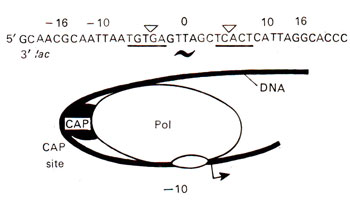
Fig. 35.12. DNA bending due to binding of CAP protein.
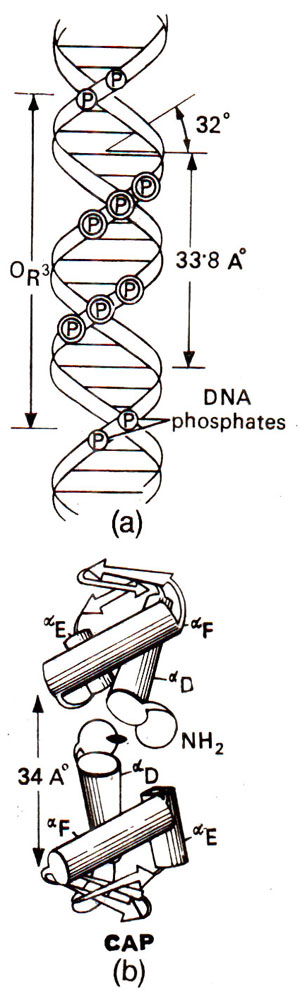
Fig. 35.13. A DNA segment and its relationship (for binding site) with CAP protein involved in regulation of gene activity. Note that CAP consists of a dimer, each monomer having three polypeptides.
Association and dissociation of lac repressor with operator. The binding of purified lac repressor protein to DNA was first characterized by a
filter binding assay. The assay is based on the principle that nitrocellulose filter retains protein or DNA complexed with protein, but not pure double stranded DNA (Fig. 35.11). Thus the DNA sequences of operator used by repressor for binding could be identified by their retention on the filter. (It was shown that repressor could not bind
; on DNA obtained from an
Oc mutant). The location of binding in
lac operon extended from - 5 to + 21 within the transcription unit, thus overlapping the right end of promoter (this overlapping does not occur in all operons). Although earlier it was believed that binding of repressor does not allow binding of RNA polymerase, now we know that both can bind simultaneously, but RNA polymerase can not initiate transcription, unless repressor is released by an inducer.

Fig. 35.11. Identification of DNA sequence to which a specific protein is bound, by nitrocellulose filter binding assay.
Repressor is a polypeptide of 360 amino acids and is divided into two regions : (i) 1-59
amino acid residues on the amino terminal end make a
long headpiece, which can be cleaved at position 51, and the fragment 1-51 is called
short headpiece, (ii) 60-360 amino acid residues on the C-terminal end make
trypsin resistant core (trypsin cleaves the repressor at amino acid no. 59). The long and short head pieces both can bind to operator independently of the core, and therefore, it is assumed that their structure does not change in association with the core. Several models have been proposed suggesting the manner of association of repressor, but all models assume that headpieces make protrusions, which help in binding.
When lactose inducer is added, repressor will dissociate from the operator. This is explained on the basis of several models : (i) Repressor is normally in equilibrium between associated and dissociated state. When the inducer binds to dissociated repressor, it is rendered incapable of association,
(ii) Inducer may bind to repressor associated with operator and changes the conformation of repressor, which allows it to get off the operator, (iii) Inducer changes the relative conformation of head and core regions of repressor, so that headpiece can not hold the DNA tightly, (iv) The repressor in tetrameric form may be in excess and remains associated at other random sites on DNA, but is not found free within the cell. Due to induction, when the repressor is dissociated from
lac operator, it is stored at other random sites and used again when needed. The effect of induction, therefore, is to change the distribution of repressor on DNA and not to release free repressor.
DNA bending during binding of CAP protein. During late 1980's and early 1990's, DNA-protein complexes have been analyzed to find if DNA geometry is altered, when a regulatory protein binds to it. It has been shown that DNA bends considerably, when the regulatory protein CAP binds to promoter site. Similar bending has been observed during binding of
zinc finger proteins or
leucine zipper proteins or TFIID (see
Expression of Gene : Protein Synthesis 2. Transcription in Prokaryotes and Eukaryotes). This bending of DNA is very important in activation of transcription. In case of CAP, bending due to CAP can be about 90°, but the protein-protein interaction between CAP and RNA polymerase leads to formation of a nucleoprotein complex showing a DNA bending of upto 180° (Fig. 35.12).

Fig. 35.12. DNA bending due to binding of CAP protein.
The CAP protein consists of 209 amino acid residues with two domains. The carboxyl terminal domain of the protein, which is involved in DNA binding, is shown in Figure 35.13. Cyclic AMP is bound only to its larger amino terminal domain. Each CAP molecule has three α helices (αD, αE, αF) and the distance between αF helices in a dimer is 34 Å making it possible for them to fit into the grooves of B DNA, which may take the form of Z-DNA during DNA-protein interaction. CAP has been compared with other proteins like λ
repressor and λ
cro protein, showing similarities in structural details and amino acid sequences in DNA-binding domains (Fig. 35.13). Comparisons of these three proteins (CAP, λRep and λCro) revealed a conserved recognition motif consisting of an α-helix, a turn and second α-helix. These are therefore described as
helix-turn-helix or
HTH proteins. (Consult
Expression of Gene : Protein Synthesis 2. Transcription in Prokaryotes and Eukaryotes).

Fig. 35.13. A DNA segment and its relationship (for binding site) with CAP protein involved in regulation of gene activity. Note that CAP consists of a dimer, each monomer having three polypeptides.


















