In
Mutations : 2. Biochemical Level (Biochemical and Microbial Genetics), we described the synthesis of amino acid tryptophan in bacteria and
Neurospora, while discussing the biochemical mutants. The five structural genes involved in the synthesis of tryptophan form an operon with an operator. These five genes are arranged in a series and code for three enzymes needed for tryptophan synthesis (Fig. 35.14). Transcription starts at the promoter site on the left of the structural genes. A repressor is coded by a gene
trp R, which is not linked with the operon. The repressor has low specificity for the operator and thus remains inactive, but can become active due to association with tryptophan which functions as a co-repressor (Fig. 35.15). The repressor functions as a tetramer of four identical subunits of 12,000 daltons. The level of tryptophan, besides working as a co-repressor, also functions through
feedback inhibition (see later in this section), where tryptophan combines with the first enzyme of tryptophan synthetic pathway and inhibits its activity through allosteric modifications (Fig. 35.16 and 35.17).
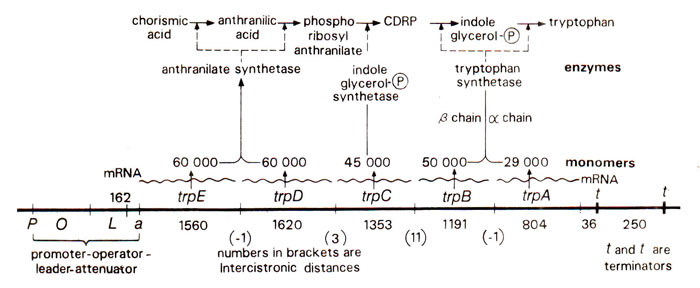
Fig. 35.14. Tryptophan operon with its five genes, the promoter-operator leader-attenuator region and the corresponding biosynthetic pathway. (CDRP carboxy phenylamino deoxyribulose phosphate).
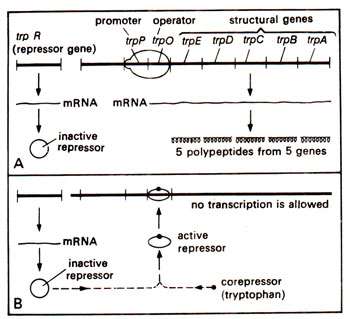
Fig. 35.15. Tryptophan operon, showing transcription in tne absence of corepressor (tryptophan) (as shown in A) and the prevention of transcription due to addition of corepressor, which activates the inactive repressor (as shown in B).
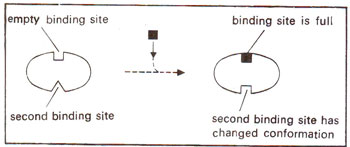
Fig. 35.16. The change in conformation of the second binding site in an allosteric protein (monomer) when the first site is occupied.

Fig. 35.17. The change in conformation of all the empty sites in an allosteric protein, when one site is occupied by a slow reaction leading to quick occupation of these empty sites.
It is thus obvious that the synthesis of tryptophan is self regulatory, since tryptophan, when present in plenty, works firstly by feed back inhibition, and secondly through co-repression to stop further synthesis of tryptophan. The major difference between lactose operon and tryptophan operon is that tryptophan increases the affinity of repressor protein for the operator. In tryptophan operon there is yet another mechanism for regulation of tryptophan synthesis through leader sequences and attenuation, which is discussed later in this section.
Tryptophan (trp) repressor controls three sets of genes
The
lac repressor acts only on
lac operon with
ZYA cluster of genes. However, a repressor can control several groups of genes dispersed in the genome.
For instance,
trp repressor controls following three sets of unlinked genes : (i)
trp EDBCA meant for tryptophan synthesis enzymes; (ii)
aroH gene coding for an enzyme needed for common pathway of aromatic amino acid biosynthesis, and (iii)
trp R regulator gene (repressed by its own gene product =
autogenous control). A related operator sequence (27bp long) is present at each of these three genes/ gene clusters, although the location of this sequence relative to start point may differ (as shown earlier). The operator may actually lie within the promoter (e.g.
trpP locus), or downstream (e.g.
lac)or upstream (e.g.
gal)
.








