Excision repair systems in E. coli
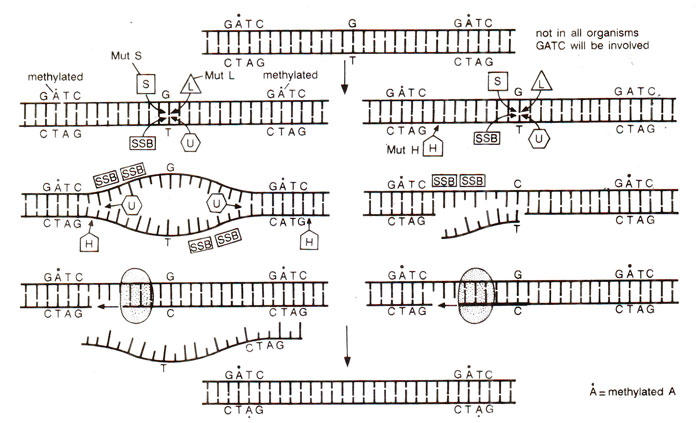
Fig. 26.34. Two models for the mechanism of mismatch repair. In both models proteins called MutL and MutS interact with mismatch site (G-T) and a protein called MutH cleaves the newly synthesized strand. The repair apparatus distinguishes the parental strand from the new one by means of methyl groups (black dots) within the parental GA-TC sequences. The strands surrounding the mismatch are separated with the help of protein MutU and stabilized by SSB. (For details of two models, see text).
Excision repair involves different lengths of DNA and is described as (i) very short patch repair (VSP), (ii) short patch repair and (iii) long patch repair. While VSP deals with single base mismatches, the other two deal with more extensive damages, which are repaired through uvr genes. (uvr A, B, C), coding for components of a repair endonuclease. Another enzyme uvr D is also needed for helicase activity. The mechanism involved in VSP is dealt in the following section under mismatch repair.
When there is mismatch in a base pair as in GC→GT, then theoretically it may repair to give rise to either wild type (GC) or to a mutant type (AT). Therefore, the repair system has to distinguish between old and new strands and repair only the new strand to restore the wild type. This is done by VSP system and requires four proteins, namely Mut L, Mut S, Mut U and Mut H coded in E. coli respectively by genes mut L, mutS, mut U and mut H. These mut genes are mutator loci, where a mutation leads to increased frequency of spontaneous mutations. It has been shown that a methylase (dam) coded by dam gene, brings about methylation in adenine of sequence GATC on both strands of DNA. Following replication, one strand remains methylated (only at A of GATC sequence) and other remains unmethylated till methylase acts on this new strand to bring about methylation. During this transition period, the unmethylated GATC allows recognition of mismatch by Mut L, and binding of Mut S to mismatch. Mut U helps in unwinding the single strands stabilised by SSB protein and Mut H cleaves the newly synthesized strand. Two models for repair have been suggested which are shown in Figure 26.34. In one model Mut H cleaves DNA at two GATC sequences flanking the mismatch, while in the other model the cleavage occurs on one side of mismatch at GATC sequence and on the other side at the mismatch itself.

Fig. 26.34. Two models for the mechanism of mismatch repair. In both models proteins called MutL and MutS interact with mismatch site (G-T) and a protein called MutH cleaves the newly synthesized strand. The repair apparatus distinguishes the parental strand from the new one by means of methyl groups (black dots) within the parental GA-TC sequences. The strands surrounding the mismatch are separated with the help of protein MutU and stabilized by SSB. (For details of two models, see text).




