In early 1960's, it was shown that some RNA viruses (e.g. Rous Sarcome Virus = RSV), after infecting the host, give rise to DNA as an intermediate step in their multiplication. This DNA then gives rise to (i) RNA for assembly of viral genomes and (ii) mRNA for synthesis of virus specific proteins. Not all RNA viruses multiply through DNA; rather many of them may actually synthesize RNA directly on RNA template as in case of influenza virus or polio virus (Fig. 26.33). The evidence for this intermediate synthesis of DNA, often described as
DNA provirus hypothesis, was initially available as follows : (i)
actinomycin D (it inhibits synthesis of RNA on DNA but not on RNA template), when added to cultured cells producing RSV, inhibited production of all RNA; (ii) when soon after inoculation with RSV, synthesis of DNA in inoculated cells was inhibited by antibiotics like
amethopterin and
fluorodeoxyuridine, no infection occurred; (iii) when cells exposed to RSV were grown in presence of
bromodeoxyuridine (its incorporation renders the newly synthesized DNA sensitive to light) and then exposed to light, no infection occurred; (iv) DNA extracted from infected cells, when denatured and hybridized with RNA, double stranded
DNA-RNA hybrid molecules could be obtained. It was also shown that the enzyme responsible for synthesis of DNA on RNA template was not freshly synthesized on infection, but was present within the virus particles (RSV).
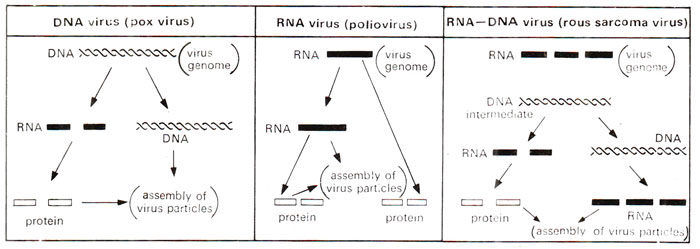
Fig. 26.33. Three kinds of viruses, grouped on the basis of genetic material and mode of replication (a) DNA viruses, (b) RNA viruses and (c) RNA:DNA viruses.
The above information on RNA directed DNA synthesis was available in 1960's but no enzyme for this activity could be discovered. In 1970, for the first time, disrupted virus particles from
RSV (used by
H. Temin) and
MLV (mouse leukaemia virus; used by
D. Baltimore) were shown to undertake DNA synthesis, utilizing dNTPs (one of them made radioactive), suggesting that an enzyme existed in the core of virus particle, which could stimulate RNA-directed DNA synthesis.
For this work later, both
Howard Temin and
David Baltimore were awarded Nobel Prize. The enzyme was initially called 'RNA dependent DNA polymerase', but was later called 'RNA directed DNA polymerase', since the purified enzyme was capable of utilizing a variety of templates including synthetic and natural DNA, RNA and RNA-DNA hybrids. The enzyme subsequently became popularly known as
reverse transcriptase. This enzyme resembles other DNA polymerases and was detected in human tumour cells, initially suggesting its role in cancerous growth of cells. However, later this enzyme was found in the normal cells also, thus reducing the initial excitement about this enzyme being the cause of cancer in humans.
Although the discovery of RNA directed DNA polymerase or reverse transcriptase did not help in providing a solution to the problem of human cancer, the enzyme is now being extensively utilized for molecular biology studies. The most important of these uses is the synthesis of complementary DNA (cDNA) from mRNA, leading to the preparation of cDNA libraries for the study of genes expressed in specific tissues. A large number of such cDNA libraries have been prepared in a variety of organisms, both plants and animals, and have facilitated isolation of genes (for more details, consult
Genetic Engineering and Biotechnology 1. Recombinant DNA and PCR (Cloning and Amplification of DNA)).

