Enumeration of Microorganisms
Introduction (Plate Count)The laboratory microbiologist often has to determine the number of bacteria in a given sample, as well as having to compare the amount of bacterial growth under various conditions. Enumeration of microorganisms is especially important in dairy microbiology, food microbiology, and water microbiology. Since the enumeration of microorganisms involves the use of extremely small dilutions and extremely large numbers of cells, scientific notation is routinely used in calculations.
The number of bacteria in a given sample is usually too great to be counted directly. However, if the sample is serially diluted and then plated out on an agar surface, single isolated bacteria can form visible isolated colonies.
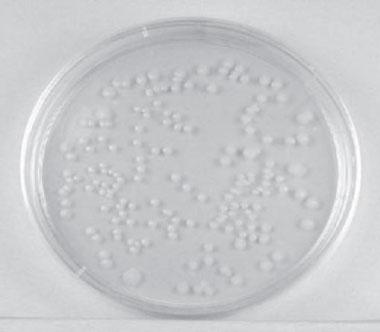 |
Figure 26 Single isolated colonies obtained during the plate count. |
The number of colonies can be used as a measure of the number of viable (living) cells in that known dilution. owever, keep in mind that if the organism normally forms multiple cell arrangements, such as chains, the colony-forming unit ay consist of a chain of bacteria rather than a single bacterium. In addition, some of the bacteria may be clumped ogether. Therefore, when doing the plate count technique, we generally say we are determining the number of colony-forming units (CFUs) in that known dilution. By extrapolation, this number can in turn be used to calculate the number of CFUs in the original sample.
Normally, the bacterial sample is diluted by factors of 10 and plated on agar. After incubation, the number of colonies on a dilution plate showing between 30 and 300 colonies is determined. A plate having 30–300 colonies is chosen, because this range is considered statistically significant. If there are less than 30 colonies on the plate, small errors in dilution technique or the presence of a few contaminants will have a drastic effect on the final count. Likewise, if there are more than 300 colonies on the plate, there will be poor isolation and colonies will have grown together.
Generally, one wants to determine the number of CFUs per milliliter (mL) of sample. To find this, the number of colonies (on a plate having 30–300 colonies) is multiplied by the number of times the original mL of bacteria were diluted (the dilution factor of the plate counted). For example, if a plate containing a 1/1,000,000 dilution of the original mL of sample shows 150 colonies, then 150 represents 1/1,000,000 the number of CFUs present in the original mL. Therefore, the number of CFUs per mL in the original sample is found by multiplying 150 × 1,000,000, as shown in the formula below:
Number of CFUs per ml of sample = number of colonies (30–300 plate) × the dilution factor of the plate counted
In the case of the example above, 150 × 1,000,000 = 150,000,000 CFUs per mL.
For a more accurate count, it is advisable to plate each dilution in duplicate or triplicate and then find an average count.
Direct Microscopic Method (Total Cell Count)
In the direct microscopic count, a counting chamber consisting of a ruled slide and a coverslip is employed. It is constructed in such a manner that a known volume is delimited by the coverslip, slide, and ruled lines. The number of bacteria in a small known volume is directly counted microscopically and the number of bacteria in the larger original sample is determined by extrapolation.
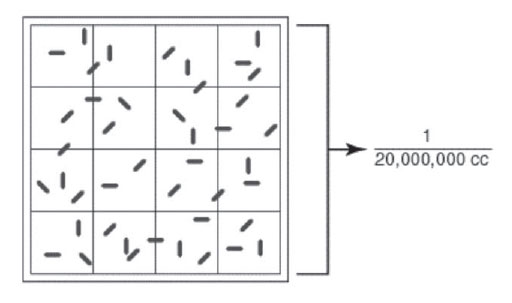 |
Figure 27 Large double-lined square of a Petroff-Hausser counter. |
 |
Figure 28 Petroff-Hausser Counter as seen through a microscope. |
The double-lined “square” holding 1/20,000,000 cc is shown by the bracket. The arrow shows a bacterium. The square holds a volume of 1/20,000,000 of a cubic centimeter. Using a microscope, the bacteria in the square are counted. For example, has squares 1/20 of a millimeter (mm) by 1/20 of a mm and is 1/50 of a mm deep. The volume of 1 square, therefore, is 1/20,000 of a cubic mm or 1/20,000,000 of a cubic centimeter (cc). The normal procedure is to count the number of bacteria in 5 large double-lined squares and divide by 5 to get the average number of acteria per large square. This number is then multiplied by 20,000,000, since the square holds a volume of 1/20,000,000 cc, to find the total number of organisms per cc in the original sample.
If the bacteria are diluted (such as by mixing with dye) before being placed in the counting chamber, then this dilution must also be considered in the final calculations.
The formula used for the direct microscopic count is:
| number of bacteria per cc = | the average number of bacteria per large double-lined square × the dilution factor of the large square (20,000,000) × the dilution factor of any dilutions made prior to placing the sample in the counting chamber, e.g., mixing the bacteria with dye. |
Turbidity
When we mix the bacteria growing in a liquid medium, the culture appears turbid. This is because a bacterial culture acts as a colloidal suspension that blocks and reflects light passing through the culture. Within limits, the light absorbed by the bacterial suspension will be directly proportional to the concentration of cells in the culture. By measuring the amount of light absorbed by a bacterial suspension, one can estimate and compare the number of bacteria present.
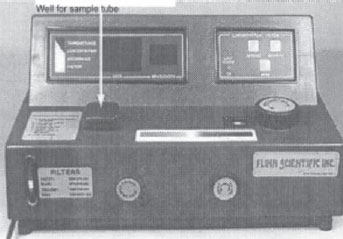 |
Figure 29 A spectrophotometer. |
The instrument used to measure turbidity is a spectrophotometer. It consists of a light source, a filter that allows only a single wavelength of light to pass through, the sample tube containing the bacterial suspension, and a photocell that compares the amount of light coming through the tube with the total light entering the tube.
The ability of the culture to block the light can be expressed as either percentage of light transmitted through the tube or amount of light absorbed in the tube.
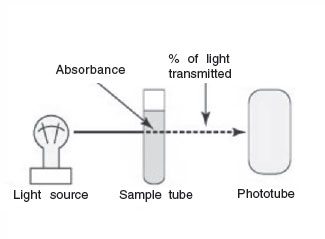 |
Figure 30 A spectrophotometer. |
The percentage of light transmitted is inversely proportional to the bacterial concentration. (The greater the percent transmittance, the lower the number of bacteria.) The absorbance (or optical density) is directly proportional to the cell concentration. (The greater the absorbance, the greater the number of bacteria.) Turbidimetric measurement is often correlated with some other method of cell count, such as the direct microscopic method or the plate count. In this way, turbidity can be used as an indirect measurement of the cell count. For example:
- Several dilutions can be made of a bacterial stock.
- A Petroff-Hausser counter can then be used to perform a direct microscopic count on each dilution.
- Then, a spectrophotometer can be used to measure the absorbance of each dilution tube.
- A standard curve comparing absorbance to the number of bacteria can be
made by plotting absorbance versus the number of bacteria per cc.
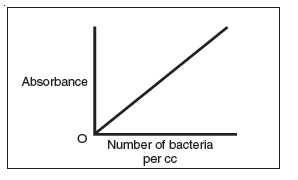 Figure 31 A standard curve plotting the number of bacteria per cc versus absorbance.
Figure 31 A standard curve plotting the number of bacteria per cc versus absorbance.
- Once the standard curve is completed, any dilution tube of that organism
can be placed in a spectrophotometer and its absorbance read. Once the
absorbance is determined, the standard curve can be used to determine the
corresponding number of bacteria per cc.
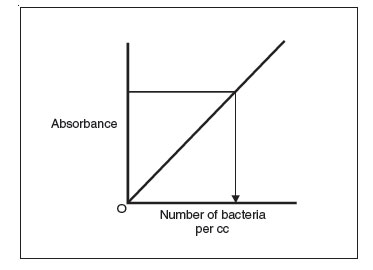 Figure 32 Using a standard curve to determine the number of bacteria per cc in a sample by measuring the sample’s absorbance.
Figure 32 Using a standard curve to determine the number of bacteria per cc in a sample by measuring the sample’s absorbance.
Materials: 6 tubes each containing 9.0 mL of sterile saline, 3 plates of trypticase soy agar, 2 sterile 1.0-mL pipettes, pipette filler, turntable, bent glass rod, dish of alcohol.
Organism: Trypticase soy broth culture of Escherichia coli.
Procedure
Plate Count
- Take 6 dilution tubes, each containing 9.0 mL of sterile saline. Aseptically
dilute 1.0 mL of a sample of E. coli, as shown in and described as follows:
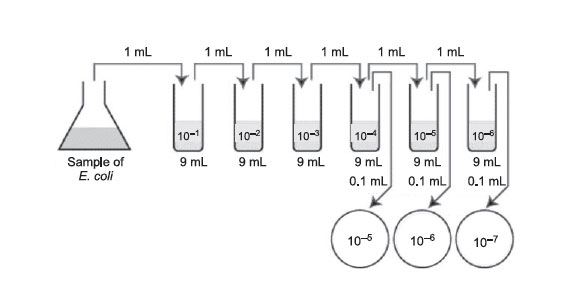 Figure 33 Plate count dilution procedure.
Figure 33 Plate count dilution procedure.
- Remove a sterile 1.0-mL pipette from the bag. Do not touch the portion of the pipette tips that will go into the tubes and do not lay the pipette down. From the tip of the pipette to the “0” line is 1 mL; each numbered division (0.1, 0.2, etc.) represents 0.1 mL.
- Insert the cotton-tipped end of the pipette into a blue 2-mL pipette filler.
- Flame the sample flask, insert the pipette to the bottom of the flask, and withdraw 1.0 mL (up to the “0” line of the sample) by turning the filler knob towards you. Draw the sample up slowly so that it isn’t accidentally drawn into the filler itself. Reflame and cap the sample.
- Flame the first dilution tube and dispense the 1.0 mL of sample into the tube by turning the filler knob away from you. Draw the liquid up and down in the pipette several times to rinse the pipette and help mix. Reflame and cap the tube.
- Mix the tube thoroughly by either holding the tube in one hand and vigorously tapping the bottom with the other hand or by using a vortex mixer. This is to assure an even distribution of the bacteria throughout the liquid.
- Using the same procedure, aseptically withdraw 1.0 mL from the first
dilution tube and dispense into the second dilution tube. Continue
doing this from tube to tube as shown in until the dilution is completed.
Discard the pipette in the biowaste disposal containers at the front of
the room and under the hood.
These pipetting and mixing techniques will be demonstrated by your
instructor.
- Using a new 1.0-mL pipette, aseptically transfer 0.1 mL from each of the
last 3 dilution tubes onto the surface of the corresponding plates of trypticase
soy agar as shown in figure. Note that since only 0.1 mL of the bacterial
dilution (rather than the desired 1.0 mL) is placed on the plate, the bacterial
dilution on the plate is 1/10 the dilution of the tube from which it came.
Using a turntable and sterile bent glass rod, immediately spread the solution
over the surface of the plates as follows:
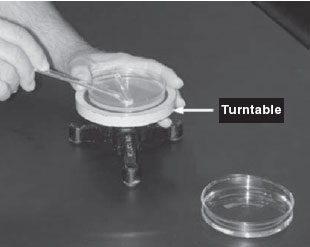 Figure 34 Using a bent glass rod and a turntable to spread a bacterial sample.
Figure 34 Using a bent glass rod and a turntable to spread a bacterial sample.
- Place the plate containing the 0.1 mL of dilution on a turntable.
- Sterilize the glass rod by dipping the bent portion in a dish of alcohol and igniting the alcohol with the flame from your burner. Let the flame burn out.
- Place the bent portion of the glass rod on the agar surface and spin the turntable for about 30 seconds to distribute the 0.1 mL of dilution evenly over the entire agar surface.
- Replace the lid and resterilize the glass rod with alcohol and flaming.
- Repeat for each plate.
- Discard the pipette in the biowaste disposal containers at the front of the room and under the hood.
- Incubate the 3 agar plates upside down at 37°C until the next lab period. Place the used dilution tubes in the disposal baskets in the hood.
Direct Microscopic Method
- Pipette 1.0 mL of the sample of E. coli into a tube containing 1.0 mL of the dye methylene blue. This produces a ½ dilution of the sample.
- Using a Pasteur pipette, fill the chamber of a Petroff-Hausser counting chamber with this ½ dilution. Turntable
- Place a coverslip over the chamber and focus on the squares using 400X (40X objective).
- Count the number of bacteria in 5 large double-lined squares. For those organisms on the lines, count those on the left and upper lines, but not those on the right and lower lines. Divide this total number by 5 to find the average number of bacteria per large square.
- Calculate the number of bacteria per cc as follows:
Number of bacteria per cc = the average number of bacteria per large
square × the dilution factor of the large
square (20,000,000) × the dilution factor of
any dilutions made prior to placing the
sample in the counting chamber, such as
mixing it with dye (2 in this case).
Your instructor will set up a spectrophotometer demonstration illustrating that as the number of bacteria in a broth culture increases, the absorbance increases (or the percent of light transmitted decreases).
Results
Plate Count
- Choose a plate that appears to have between 30 and 300 colonies.
- Sample 1/100,000 dilution plate
- Sample 1/1,000,000 dilution plate
- Sample 1/10,000,000 dilution plate.
- Count the exact number of colonies on that plate using the colony counter (as demonstrated by your instructor).
- Calculate the number of CFUs per mL of original sample as follows:
Number of CFUs per mL of sample = Number of colonies (30–300 plate) × the dilution factor of the plate counted
____________ = Number of colonies
____________ = Dilution factor of plate counted
____________ = Number of CFUs per mL.
- Record your results on the blackboard.
Observe the demonstration of the Petroff-Hausser counting chamber.
Turbidity
Observe your instructor’s demonstration of the spectrophotometer.
Performance Objectives
Discussion
- Provide the formula for determining the number of CFUs per mL of sample when using the plate count technique.
- When given a diagram of a plate count dilution and the number of colonies
on the resulting plates, choose the correct plate for counting, determine the
dilution factor of that plate, and calculate the number of CFUs per mL in
the original sample.
- Plate count practice problems- A sample of E.coli is diluted according to the above diagram. The number of colonies that grew is indicated on the petri plates. How many CFUs are there per mL in the original sample?
- The correct plate is the one having 63 colonies on the 1/1,000,000 or 10–6 dilution.
- Multiply 63 by the dilution factor of that plate.
- Since the dilution factor is 1/1,000,000 or 10–6, the dilution factor or inverse is 1,000,000, or 106.
- 63 × 1,000,000 = 63,000,000 CFUs per ml (6.3 × 107 in scientific notation.)
- A sample of E.coli is diluted according to the above diagram. The number
of colonies that grew is indicated on the petri plates. How many CFUs are
there per mL in the original sample?
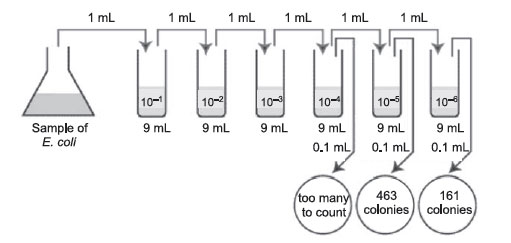 Figure 37 Plate count: Practice problem B.
Figure 37 Plate count: Practice problem B.
The correct dilutions are shown on the tubes and plates above.
The formula to be used is:
- First choose the correct plate to count, that is, one with between 30 and
300 colonies.
- The correct plate is the one having 161 colonies on the 1/1,000,000 or 10–6 dilution.
- Multiply 161 by the dilution factor of that plate.
- Since the dilution factor is 1/1,000,000 or 10–6, the dilution factor or inverse is 1,000,000, or 106.
- 161 × 1,000,000 =161,000,000 CFUs per mL (1.61 × 108 in scientific notation.)
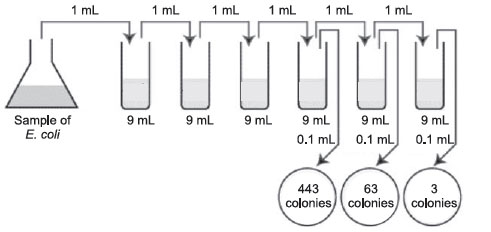 Figure 35 Plate count: Practice problem A.
Figure 35 Plate count: Practice problem A.
The correct dilutions are shown on the tubes and plates above. The formula to be used is:
Number of CFUs per mL of sample = Number of colonies (30–300 plate) × the dilution factor of the plate counted
- First choose the correct plate to count, that is, one with between 30 and 300 colonies.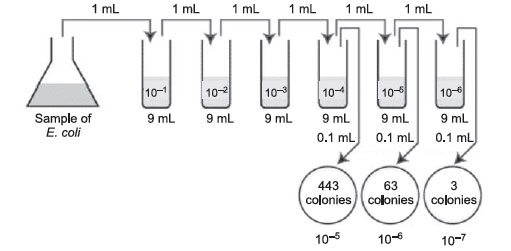 Figure 36 Plate count: Answer to practice problem A.
Figure 36 Plate count: Answer to practice problem A.Number of CFUs per mL of sample = Number of colonies (30–300 plate) × the dilution factor of the plate counted
- Explain the principle behind the direct microscopic method of enumeration.
- Provide the formula for determining the number of bacteria per cc of sample when using the direct microscopic method of enumeration.
- When given the total number of bacteria counted in a Petroff-Hausser chamber, the total number of large squares counted, and the dilution of the bacteria placed in the chamber, calculate the total number of bacteria per cc in the original sample.
- Describe the function of a spectrophotometer.
- Explain the relationship between absorbance (optical density) and the number of bacteria in a broth sample.
- Explain the relationship between percent of light transmitted and the number of bacteria in a broth sample.
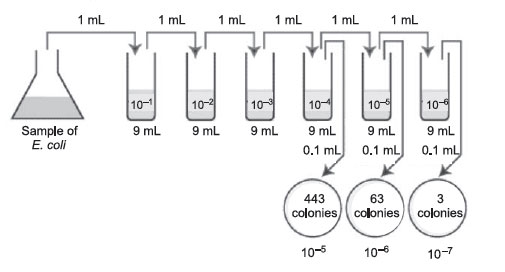 |
Figure 38 Plate count: Answer to practice problem B. |
Perform a serial dilution of a bacterial sample, according to instructions in the lab manual, and plate out samples of each dilution using the spin-plate technique.
Results
Using a colony counter, count the number of colonies on a plate showing between 30 and 300 colonies and, by knowing the dilution of this plate, calculate the number of CFUs per mL in the original sample.




