Isolation and Identification of Staphylococci
The laboratory diagnosis of staphylococcal disease is made by identifying the organism (usually S. aureuss) in a clinical specimen representing the site of infection (pus from a skin lesion, sputum when pneumonia is suspected, urine, spinal fluid, or blood). It should be remembered that either S. aureussor S. epidermidis may be harmlessly present on superficial tissues. Special care must be taken not to contaminate the specimen with normal flora, and laboratory results must be interpreted in the light of the patient’s clinical symptoms. The principal features by which staphylococci are recognized and distinguished in the laboratory include their microscopic morphology, colonial appearance on blood agar (especially hemolytic activity), coagulase activity(see colorplate 26), reaction to the carbohydrate mannitol, and susceptibility to the antimicrobial agent novobiocin (see colorplate 27)latex agglutination test is available for identifying S. aureus from characteristic colonies growing on agar media. The antibodycoated latex beads react with two surface proteins typically found on S. aureus strains. One is a type of coagulase bound to the agulase bound to the staphylococcal surface, and the other is a surface protein known as protein A (see fig. 20.1). The species are indistinguishaOn blood agar, S. aureus usually displays a light to golden yellow pigment (hence, its name), whereas S. epidermidis whereas S. epidermidis has a white pigment and S. saprophyticus either a bright yellow or white pigment. However, pigmentation ischaracteristic. On blood agar, S. aureus is usually, but not always, beta-hemolytic; S. epidermidis and S. saprophyticus are almost always nonhemolytic. S. aureus is, by definition, coagulase positive; S. epidermidis and S. saprophyticus are coagulase negative. Other coagulase negative. Other Staphylococcus species that are found on skin but seldom cause disease are also coagulase negative. As a group, these species, along with S. epidermidis and S. saprophyticus are referred to as coagulase-negative staphylococci. S. aureus is further distinguished by its ability to ferment mannitol, and S. saprophyticus by its resistance to low concentrations of novobiocin (see colorplate 27).Since specimens from the mucous membranes or skin may contain a mixed normal flora as well as the pathogenic staphylococci being sought, the use of a selective, differential medium in the primary isolation battery can be very helpful (see table 16.1). Mannitol salt agar is such a medium. It contains a high concentration of salt that inhibits gram-positive cocci other than staphylococci and many other organisms as well. It also contains mannitol and an indicator to differentiate S. aureus strains from coagulase-negative staphylococci growing on it. A blood agar plate is also essential for demonstrating hemolytic organisms. Since some streptococci, as well as many strains of S. aureus, are beta-hemolytic, they can be distinguished promptly. Aside from microscopic morphology, the simplest, most rapid distinction can be made with the catalase test, for all streptococci are catalase negative, whereas all staphylococci are catalase positive.
Staphylococcus aureus is carried by a large segment of the population as a member of the normal flora. It causes disease primarily in individuals with lowered resistance, particularly patients in hospitals. In the hospital, S. aureus is a major cause of nosocomial infections transmitted from hospital personnel or the environment.
The problem is compounded by the fact that many “hospital” strains of staphylococci are resistant to the useful antimicrobial agents. All personnel involved in patient care should be knowledgeable of transmission routes
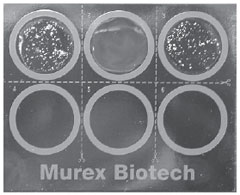 |
| Figure 20.1 A rapid latex agglutinationtest for identifying Staphylococcus aureus. The top left and center wells are the positive and negative controls, respectively. The top right well is the positive reaction of the patient’s isolate. |
and carefully follow strict procedures designed to prevent nosocomial infection. In the experiments that follow, you will be seeing and handling staphylococcal cultures. Use your knowledge of aseptic technique and make certain that you do not carry staphylococci out of the laboratory as new additions to the flora of your hands or clothes. Keep your hands scrupulously clean. If you have any minor cuts or scratches or other injury to your hands, they should be protected. While in the laboratory, keep your hands and implements with which you are working away from your mouth and face.
| Purpose | To isolate and identify staphylococci |
| Materials | Blood agar plate (BAP) Mannitol salt agar plate (MSA) Tubed plasma (0.5-ml aliquots) Novobiocin disks (5 µg) Sterile 1.0-ml pipettes Pipette bulb or other aspiration device Latex agglutination kit for Staphylococcus aureus 24-hour broth cultures of Staphylococcus epidermidis, Staphylococcus aureus, Staphylococcus saprophyticus, and Escherichia coli |
Procedures
- With your marking pencil, divide the bottom of a BAP and an MSA plate into four segments each.
- Using the grown broth cultures, inoculate one section of each plate with S. aureus, one with S. epidermidis, one with S. saprophyticus, and one with E. coli. Streak each section carefully, remaining within the assigned space.
- With heated and cooled forceps, pick up a novobiocin disk, place it in the center of one of the streaked areas of the BAP, and press it gently onto the agar with the forcep tips.
- Repeat step 3 for the remaining three organisms on the streaked BAP.
- Place the plates in the 35°C incubator for 24 hours.
- Perform a coagulase test on each of the three Staphylococcus broth cultures as follows:
- Using a sterile pipette, measure 0.1 ml of the S. epidermidis broth culture with the aspiration device. Transfer this inoculum to a tube of plasma. Discard the pipette in disinfectant. Label the tube.
- Inoculate a second and third tube of plasma with 0.1 ml of the S. aureus and S. saprophyticus broth cultures, respectively, as in step 6a.
- Place all inoculated plasma tubes in the 35°C incubator. After 30 minutes, remove and examine them (close the incubator door while you read them). Hold the tubes in a semihorizontal position to see whether the plasma in the tube is beginning to clot into a solid mass. If so, make a record of the tube showing coagulase activity. Return unclotted tubes to the incubator.
- Repeat procedure 6c every 30 minutes for 4 hours, if necessary.
- After 24 hours of incubation of the plate cultures prepared in procedures 1 and 2, examine and record colonial morphology. Make Gram stains of each culture on the BAP and record microscopic morphology. Measure and record the diameter of the zone of inhibition around the novobiocin disks. A zone size greater than 12 mm in diameter is considered susceptible.
- Following the instructor’s directions, place one drop of the latex agglutination reagent onto each of two circles on the card provided. With the special stick contained in the kit or a sterile inoculating loop, pick up several colonies of S. aureus from the blood agar plate you inoculated at the previous laboratory session. Emulsify the colonies in the latex reagent, being careful not to scratch the card. Repeat this procedure with colonies of S. epidermidis. Do not use colonies from the mannitol agar plate as these are difficult to emulsify.
- Rotate the card gently for 20 seconds, observing the circles for a clearly visible clumping of the latex particles and a clearing of the milky background (see fig. 20.1). This reaction signifies a positive test. Record the results in the chart, then dispose of the reaction card in the disinfectant provided.
Results
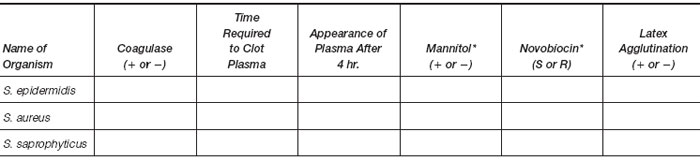
- Table of plate culture results.
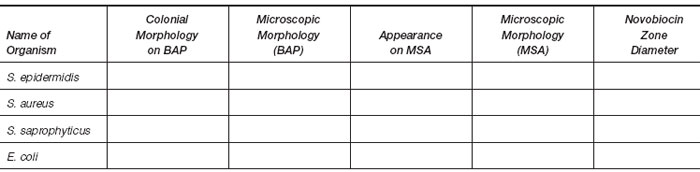
- Results of identification tests.
*Your interpretation of results from MSA and novobiocin plates.