Laboratory Diagnosis of a Sore Throat
| Purpose | To identify bacterial species in a simulated clinical throat culture as quickly as possible |
| Materials | Swab in a tube of broth, accompanied by a laboratory request for culture
Forceps Bacitracin disks (A disks) Tubes containing 0.4 ml streptococcal extraction enzyme or prepared extract Capillary pipettes Latex test kit for serological typing |
Procedures
- Using the swab in the “specimen” tube, inoculate a small area of the blood agar plate. Discard the swab in disinfectant solution. With a sterilized inoculating loop, streak the remainder of the plate to obtain isolated colonies. After you have completed the streaking step, make a few shallow cuts with your loop in the area of the original inoculum.
-
Incubate the plate at 35°C for 24 hours.
-
After the plate has incubated, examine it carefully for the presence of hemolysis and record the type of hemolysis you see on the laboratory work card. Record the colonial morphology and make Gram stains of different colony types.
-
On the basis of your findings, record on the Microbiology Laboratory Report the preliminary result that you will give to the “physician” when he or she calls for a report.
-
With your sterilized inoculating loop, pick up a few colonies that appear to produce beta-hemolysis. Streak the inoculum heavily on a portion of a blood agar plate. Using heated and cooled forceps, place a bacitracin disk on the area of heavy inoculum.
-
Incubate the plate for 24 hours at 35°C and record the result on the laboratory work card.
-
If the instructor has not prepared an extract of beta-hemolytic colonies grown from the patient’s throat specimen, follow steps 8 and 9.
-
With your sterile loop, make a light suspension of “suspicious” beta-hemolytic colonies in 0.4 ml of extraction enzyme. Five or six colonies should be sufficient.
-
Place the suspension in a 37°C water bath or in a beaker of water warmed to 37°C in an incubator. After 5 minutes, shake the tube and continue incubating for no less than 10 minutes and up to one hour.
-
Following the manufacturer’s directions, mix one drop of group A latex reagent and one drop of group B latex reagent each with a drop of your extract on a glass slide or special reaction card provided. Rock the slide back and forth for at least one minute looking for the formation of agglutinated latex particles and a clearing of the background (see fig. 21.1).
-
If agglutination is present, record the group (A or B) on your work card along with the final organism identification(s).
-
Complete the Microbiology Laboratory Report for the “physician.”
Results
- Laboratory work card (record of your work to be kept on file for at least two years).
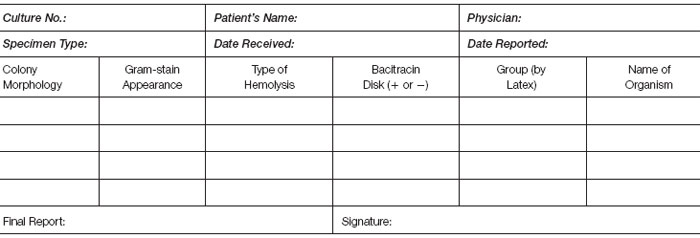
- Final laboratory report to “physician.”